Explaining pQTLS
Briana Mittleman
11/29/2018
Last updated: 2018-12-03
workflowr checks: (Click a bullet for more information)-
✔ R Markdown file: up-to-date
Great! Since the R Markdown file has been committed to the Git repository, you know the exact version of the code that produced these results.
-
✔ Environment: empty
Great job! The global environment was empty. Objects defined in the global environment can affect the analysis in your R Markdown file in unknown ways. For reproduciblity it’s best to always run the code in an empty environment.
-
✔ Seed:
set.seed(12345)The command
set.seed(12345)was run prior to running the code in the R Markdown file. Setting a seed ensures that any results that rely on randomness, e.g. subsampling or permutations, are reproducible. -
✔ Session information: recorded
Great job! Recording the operating system, R version, and package versions is critical for reproducibility.
-
Great! You are using Git for version control. Tracking code development and connecting the code version to the results is critical for reproducibility. The version displayed above was the version of the Git repository at the time these results were generated.✔ Repository version: 37ef750
Note that you need to be careful to ensure that all relevant files for the analysis have been committed to Git prior to generating the results (you can usewflow_publishorwflow_git_commit). workflowr only checks the R Markdown file, but you know if there are other scripts or data files that it depends on. Below is the status of the Git repository when the results were generated:
Note that any generated files, e.g. HTML, png, CSS, etc., are not included in this status report because it is ok for generated content to have uncommitted changes.Ignored files: Ignored: .DS_Store Ignored: .Rhistory Ignored: .Rproj.user/ Ignored: data/.DS_Store Ignored: output/.DS_Store Untracked files: Untracked: KalistoAbundance18486.txt Untracked: analysis/DirectionapaQTL.Rmd Untracked: analysis/ncbiRefSeq_sm.sort.mRNA.bed Untracked: analysis/snake.config.notes.Rmd Untracked: analysis/verifyBAM.Rmd Untracked: data/18486.genecov.txt Untracked: data/APApeaksYL.total.inbrain.bed Untracked: data/ChromHmmOverlap/ Untracked: data/GM12878.chromHMM.bed Untracked: data/GM12878.chromHMM.txt Untracked: data/LocusZoom/ Untracked: data/NuclearApaQTLs.txt Untracked: data/PeakCounts/ Untracked: data/PeaksUsed/ Untracked: data/RNAkalisto/ Untracked: data/TotalApaQTLs.txt Untracked: data/Totalpeaks_filtered_clean.bed Untracked: data/YL-SP-18486-T-combined-genecov.txt Untracked: data/YL-SP-18486-T_S9_R1_001-genecov.txt Untracked: data/apaExamp/ Untracked: data/bedgraph_peaks/ Untracked: data/bin200.5.T.nuccov.bed Untracked: data/bin200.Anuccov.bed Untracked: data/bin200.nuccov.bed Untracked: data/clean_peaks/ Untracked: data/comb_map_stats.csv Untracked: data/comb_map_stats.xlsx Untracked: data/comb_map_stats_39ind.csv Untracked: data/combined_reads_mapped_three_prime_seq.csv Untracked: data/diff_iso_trans/ Untracked: data/ensemble_to_genename.txt Untracked: data/example_gene_peakQuant/ Untracked: data/filtered_APApeaks_merged_allchrom_refseqTrans.closest2End.bed Untracked: data/filtered_APApeaks_merged_allchrom_refseqTrans.closest2End.noties.bed Untracked: data/first50lines_closest.txt Untracked: data/gencov.test.csv Untracked: data/gencov.test.txt Untracked: data/gencov_zero.test.csv Untracked: data/gencov_zero.test.txt Untracked: data/gene_cov/ Untracked: data/joined Untracked: data/leafcutter/ Untracked: data/merged_combined_YL-SP-threeprimeseq.bg Untracked: data/mol_overlap/ Untracked: data/mol_pheno/ Untracked: data/nom_QTL/ Untracked: data/nom_QTL_opp/ Untracked: data/nom_QTL_trans/ Untracked: data/nuc6up/ Untracked: data/other_qtls/ Untracked: data/pQTL_otherphen/ Untracked: data/peakPerRefSeqGene/ Untracked: data/perm_QTL/ Untracked: data/perm_QTL_opp/ Untracked: data/perm_QTL_trans/ Untracked: data/perm_QTL_trans_filt/ Untracked: data/reads_mapped_three_prime_seq.csv Untracked: data/smash.cov.results.bed Untracked: data/smash.cov.results.csv Untracked: data/smash.cov.results.txt Untracked: data/smash_testregion/ Untracked: data/ssFC200.cov.bed Untracked: data/temp.file1 Untracked: data/temp.file2 Untracked: data/temp.gencov.test.txt Untracked: data/temp.gencov_zero.test.txt Untracked: output/picard/ Untracked: output/plots/ Untracked: output/qual.fig2.pdf Unstaged changes: Modified: analysis/28ind.peak.explore.Rmd Modified: analysis/39indQC.Rmd Modified: analysis/apaQTLoverlapGWAS.Rmd Modified: analysis/cleanupdtseq.internalpriming.Rmd Modified: analysis/coloc_apaQTLs_protQTLs.Rmd Modified: analysis/dif.iso.usage.leafcutter.Rmd Modified: analysis/diff_iso_pipeline.Rmd Modified: analysis/explore.filters.Rmd Modified: analysis/flash2mash.Rmd Modified: analysis/overlapMolQTL.Rmd Modified: analysis/overlap_qtls.Rmd Modified: analysis/peakOverlap_oppstrand.Rmd Modified: analysis/pheno.leaf.comb.Rmd Modified: analysis/swarmPlots_QTLs.Rmd Modified: analysis/test.max2.Rmd Modified: code/Snakefile
Expand here to see past versions:
In this analysis I want to look at how well the eQTLs and apaQTLs an explain pQTLs. I will use linear models on the effect sizes.
Libraries
library(workflowr)This is workflowr version 1.1.1
Run ?workflowr for help getting startedlibrary(tidyverse)── Attaching packages ───────────────────────────────────────────────────────── tidyverse 1.2.1 ──✔ ggplot2 3.0.0 ✔ purrr 0.2.5
✔ tibble 1.4.2 ✔ dplyr 0.7.6
✔ tidyr 0.8.1 ✔ stringr 1.3.1
✔ readr 1.1.1 ✔ forcats 0.3.0── Conflicts ──────────────────────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()library(reshape2)
Attaching package: 'reshape2'The following object is masked from 'package:tidyr':
smithslibrary(broom)
library(cowplot)
Attaching package: 'cowplot'The following object is masked from 'package:ggplot2':
ggsaveInput the pQTLs (10% FDR) and gene names
geneNames=read.table("../data/ensemble_to_genename.txt", sep="\t", header=T,stringsAsFactors = F)
pQTL=read.table("../data/other_qtls/fastqtl_qqnorm_prot.fixed.perm.out", col.names = c("Gene.stable.ID", "nvar", "shape1", "shape2", "dummy", "sid", "dist", "npval", "slope", "ppval", "bpval"), header = F, stringsAsFactors = F) %>% inner_join(geneNames, by="Gene.stable.ID") %>% dplyr::select("Gene.name","Gene.stable.ID", "nvar", "shape1", "shape2", "dummy", "sid", "dist", "npval", "slope", "ppval", "bpval")
pQTL$bh=p.adjust(pQTL$bpval, method="fdr")
pQTL_sig=pQTL %>% filter(-log10(bh)> 1) Example CCDC51
Start with the exanple with the highest effect size:
CCDC51- ENSG00000164051 3:48476431
I need all of the results for this snp gene pair from the total, nuclear, and RNA nominal files. I can make a python script that will take the gene name and snp and create the relevent dataframe.
files: * /project2/gilad/briana/threeprimeseq/data/nominal_APAqtl_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Nuclear_NomRes.txt
* /project2/gilad/briana/threeprimeseq/data/nominal_APAqtl_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Total_NomRes.txt
* /project2/gilad/briana/threeprimeseq/data/molecular_QTLs/nom/fastqtl_qqnorm_RNAseq_phase2.fixed.nominal.out (need the ENSG ID)
APAandRNAfromProtQTLs.py
#use this by inserting a gene, gene_ensg (from prot), and snp for the protien QTLs
def main(gene, gene_ensg,snp):
out_prot=open("/project2/gilad/briana/threeprimeseq/data/protQTL_otherphen/%s_%s_Protres.txt"%(gene, snp), "w")
out_RNA=open("/project2/gilad/briana/threeprimeseq/data/protQTL_otherphen/%s_%s_RNAres.txt"%(gene, snp), "w")
out_Total=open("/project2/gilad/briana/threeprimeseq/data/protQTL_otherphen/%s_%s_Totalres.txt"%(gene, snp), "w")
out_Nuclear=open("/project2/gilad/briana/threeprimeseq/data/protQTL_otherphen/%s_%s_Nuclearres.txt"%(gene, snp), "w")
for ln in open("/project2/gilad/briana/threeprimeseq/data/nominal_APAqtl_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Nuclear_NomRes.txt", "r"):
s=ln.split()[1]
g=ln.split()[0].split(":")[3].split("_")[0]
if g==gene:
out_Nuclear.write(ln)
for ln in open("/project2/gilad/briana/threeprimeseq/data/nominal_APAqtl_trans/filtered_APApeaks_merged_allchrom_refseqGenes_pheno_Total_NomRes.txt", "r"):
s=ln.split()[1]
g=ln.split()[0].split(":")[3].split("_")[0]
if g==gene:
out_Total.write(ln)
for ln in open("/project2/gilad/briana/threeprimeseq/data/molecular_QTLs/nom/fastqtl_qqnorm_RNAseq_phase2.fixed.nominal.out", "r"):
s=ln.split()[1]
g=ln.split()[0]
if gene_ensg in g:
out_RNA.write(ln)
for ln in open("/project2/gilad/briana/threeprimeseq/data/molecular_QTLs/nom/fastqtl_qqnorm_prot.fixed.nominal.out", "r"):
s=ln.split()[1]
g=ln.split()[0]
if gene_ensg in g:
out_prot.write(ln)
out_Total.close()
out_Nuclear.close()
out_RNA.close()
out_prot.close()
if __name__ == "__main__":
import sys
gene=sys.argv[1]
gene_ensg=sys.argv[2]
snp=sys.argv[3]
main(gene, gene_ensg, snp)
CCDC51_APAandRNAfromProtQTLs.sh
#!/bin/bash
#SBATCH --job-name=CCDC51_APAandRNAfromProtQTLs
#SBATCH --account=pi-yangili1
#SBATCH --time=24:00:00
#SBATCH --output=CCDC51_APAandRNAfromProtQTLs.out
#SBATCH --error=CCDC51_APAandRNAfromProtQTLs.err
#SBATCH --partition=broadwl
#SBATCH --mem=12G
#SBATCH --mail-type=END
module load python
python APAandRNAfromProtQTLs.py CCDC51 ENSG00000164051 3:48476431 I first want to look at the effect sizes. I am interested in understanding the directions of the effect sizes.
3:48728204. This genotype is in the genotype file twice. I am gonig to remove this snp from the analysis.
nom_names=c("gene", "snp", "dist", "pval", "slope")
CCDC51_apaTot=read.table("../data/pQTL_otherphen/CCDC51_3:48476431_Totalres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope)
CCDC51_apaTot=filter(CCDC51_apaTot, !grepl("3:48728204",snp))
CCDC51_apaNuc=read.table("../data/pQTL_otherphen/CCDC51_3:48476431_Nuclearres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope)
CCDC51_apaNuc=filter(CCDC51_apaNuc, !grepl("3:48728204",snp))
CCDC51_RNA=read.table("../data/pQTL_otherphen/CCDC51_3:48476431_RNAres.txt", col.names = nom_names, stringsAsFactors = F) %>% select(snp, slope)
CCDC51_RNA=filter(CCDC51_RNA, !grepl("3:48728204",snp))
CCDC51_Prot=read.table("../data/pQTL_otherphen/CCDC51_3:48476431_Protres.txt", col.names = nom_names, stringsAsFactors = F)%>% select(snp, slope)
CCDC51_Prot=filter(CCDC51_Prot, !grepl("3:48728204",snp))Start with RNA and protein because they are the most simple
CCDC51_ProtRNA=CCDC51_RNA %>% left_join(CCDC51_Prot, by="snp")
colnames(CCDC51_ProtRNA)=c("snp", "RNA", "Protein")Model:
CCDC51_lmProtRNA=lm(Protein ~ RNA, data=CCDC51_ProtRNA)
CCDC51_lmProtRNA_sum=glance(CCDC51_lmProtRNA)Plot this:
plot(CCDC51_ProtRNA$Protein ~ CCDC51_ProtRNA$RNA)
abline(lm(CCDC51_ProtRNA$Protein ~ CCDC51_ProtRNA$RNA))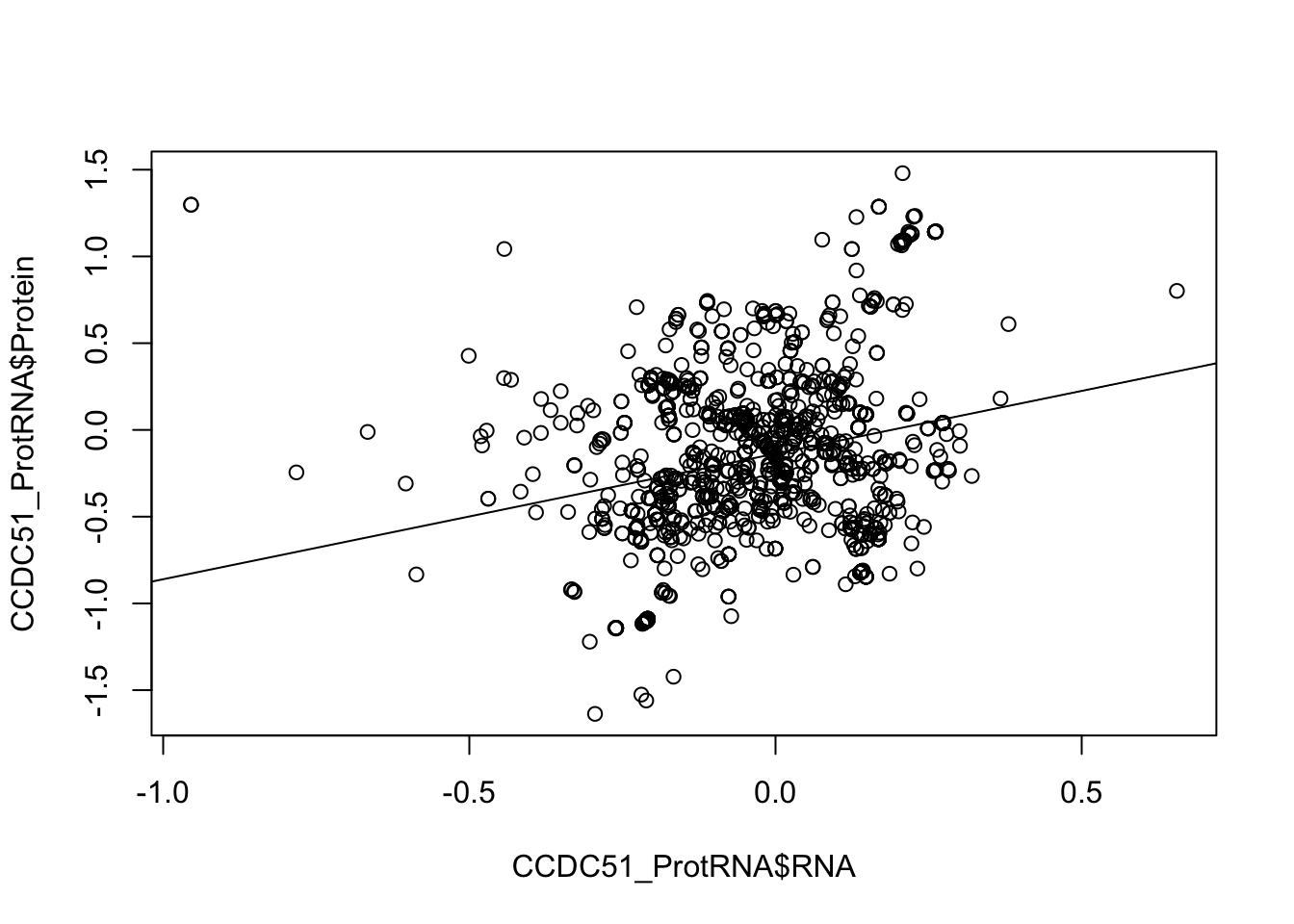 For APA I need to get a data frame that has the snps by the effect sizes for each peak.I basically want to spread this df.
For APA I need to get a data frame that has the snps by the effect sizes for each peak.I basically want to spread this df.
CCDC51_apaTot_s=spread(CCDC51_apaTot, "peak", "slope",drop = F)CCDC51_apaNuc_s=spread(CCDC51_apaNuc, "peak", "slope",drop = F)I can look at the protein ~ apaTotal.
CCDC51_apaTotProt=CCDC51_apaTot_s %>% left_join(CCDC51_Prot, by="snp")
colnames(CCDC51_apaTotProt)=c("snp", "peak216857", "peak216858", "peak216859", "peak216860", "peak216867","Protein")CCDC51_lmProtAPA=lm(Protein ~peak216857 + peak216858 + peak216859 + peak216860 + peak216867, data=CCDC51_apaTotProt)
CCDC51_lmProtAPA_sum=glance(CCDC51_lmProtAPA)Try with all of it:
CCDC51_apaTotProtRNA=CCDC51_apaTot_s %>% left_join(CCDC51_ProtRNA, by="snp")
colnames(CCDC51_apaTotProtRNA)=c("snp", "peak216857", "peak216858", "peak216859", "peak216860", "peak216867","RNA","Protein")CCDC51_lmProtAPARNA=lm(Protein ~RNA +peak216857 + peak216858 + peak216859 + peak216860 + peak216867, data=CCDC51_apaTotProtRNA)
CCDC51_lmProtAPARNA_sum=glance(CCDC51_lmProtAPARNA)
adjR2_CCDC51=cbind("CCDC51", CCDC51_lmProtRNA_sum[1,2], CCDC51_lmProtAPA_sum[1,2], CCDC51_lmProtAPARNA_sum[1,2])
colnames(adjR2_CCDC51)=c("Gene", "RNA", "APA", "RNAandAPA")Example DTD1
DTD1- ENSG00000125821 20:18607243
DTD1_APAandRNAfromProtQTLs.sh
#!/bin/bash
#SBATCH --job-name=DTD1_APAandRNAfromProtQTLs
#SBATCH --account=pi-yangili1
#SBATCH --time=24:00:00
#SBATCH --output=DTD1_APAandRNAfromProtQTLs.out
#SBATCH --error=DTD1_APAandRNAfromProtQTLs.err
#SBATCH --partition=broadwl
#SBATCH --mem=12G
#SBATCH --mail-type=END
module load python
python APAandRNAfromProtQTLs.py DTD1 ENSG00000125821 20:18607243nom_names=c("gene", "snp", "dist", "pval", "slope")
DTD1_RNA=read.table("../data/pQTL_otherphen/DTD1_20:18607243_RNAres.txt", col.names = nom_names, stringsAsFactors = F) %>% select(snp, slope) %>% filter(!duplicated(snp))
DTD1_Prot=read.table("../data/pQTL_otherphen/DTD1_20:18607243_Protres.txt", col.names = nom_names, stringsAsFactors = F)%>% select(snp, slope) %>% filter(!duplicated(snp))
#FILTER snps in the list from Rna and prot
DTD1_apaTot=read.table("../data/pQTL_otherphen/DTD1_20:18607243_Totalres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp))
DTD1_apaNuc=read.table("../data/pQTL_otherphen/DTD1_20:18607243_Nuclearres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp)) Start with RNA and protein because they are the most simple
DTD1_ProtRNA=DTD1_RNA %>% left_join(DTD1_Prot, by="snp")
colnames(DTD1_ProtRNA)=c("snp", "RNA", "Protein")Model:
DTD1_lmProtRNA=lm(Protein ~ RNA, data=DTD1_ProtRNA)
DTD1_lmProtRNA_sum=glance(DTD1_lmProtRNA)DTD1_apaTot_s=spread(DTD1_apaTot, "peak", "slope",drop = F)DTD1_apaNuc_s=spread(DTD1_apaNuc, "peak", "slope",drop = F)DTD1_apaTotProt=DTD1_apaTot_s %>% inner_join(DTD1_Prot, by="snp")
colnames(DTD1_apaTotProt)=c("snp", "peak195416", "peak195418" ,"peak195419", "peak195420", "peak195423", "peak195424","peak195425" ,"peak195426", "peak195427", "peak195428", "peak195429", "peak195430", "peak195431","peak195432", "peak195433" ,"peak195434" ,"peak195436", "peak195438", "peak195443", "peak195444" ,"peak195445", "peak195446", "peak195447", "peak195449" ,"peak195450", "peak195451", "peak195452","peak195453", "peak195454", "peak195455" ,"protein")DTD1_lmProtAPA=lm(protein ~ peak195416+ peak195418 +peak195419+peak195420+ peak195423+ peak195424+peak195425 +peak195426+ peak195427+ peak195428+ peak195429+ peak195430 + peak195431+peak195432 + peak195433+peak195434+peak195436+peak195438+peak195443+peak195444+peak195445+peak195446+peak195447+peak195449+peak195450+peak195451+peak195452+peak195453+ peak195454 +peak195455, data=DTD1_apaTotProt)
DTD1_lmProtAPA_sum=glance(DTD1_lmProtAPA)Try with all:
DTD1_apaTotProtRNA=DTD1_apaTot_s %>% inner_join(DTD1_ProtRNA, by="snp")
colnames(DTD1_apaTotProtRNA)=c("snp", "peak195416", "peak195418" ,"peak195419", "peak195420", "peak195423", "peak195424","peak195425" ,"peak195426", "peak195427", "peak195428", "peak195429", "peak195430", "peak195431","peak195432", "peak195433" ,"peak195434" ,"peak195436", "peak195438", "peak195443", "peak195444" ,"peak195445", "peak195446", "peak195447", "peak195449" ,"peak195450", "peak195451", "peak195452","peak195453", "peak195454", "peak195455" ,"RNA", "protein")DTD1_lmProtAPARNA=lm(protein ~RNA+ peak195416+ peak195418 +peak195419+peak195420+ peak195423+ peak195424+peak195425 +peak195426+ peak195427+ peak195428+ peak195429+ peak195430 + peak195431+peak195432 + peak195433+peak195434+peak195436+peak195438+peak195443+peak195444+peak195445+peak195446+peak195447+peak195449+peak195450+peak195451+peak195452+peak195453+ peak195454 +peak195455, data=DTD1_apaTotProtRNA)
DTD1_lmProtAPARNA_sum=glance(DTD1_lmProtAPARNA)
adjR2_DTD1=cbind("DTD1",DTD1_lmProtRNA_sum[1,2], DTD1_lmProtAPA_sum[1,2], DTD1_lmProtAPARNA_sum[1,2])
colnames(adjR2_DTD1)=c("Gene", "RNA", "APA", "RNAandAPA")Example DHRS7B
DHRS7B ENSG00000109016 17:21036822 DHRS7B_APAandRNAfromProtQTLs.sh
#!/bin/bash
#SBATCH --job-name=DHRS7B_APAandRNAfromProtQTLs
#SBATCH --account=pi-yangili1
#SBATCH --time=24:00:00
#SBATCH --output=DHRS7B_APAandRNAfromProtQTLs.out
#SBATCH --error=DHRS7B_APAandRNAfromProtQTLs.err
#SBATCH --partition=broadwl
#SBATCH --mem=12G
#SBATCH --mail-type=END
module load python
python APAandRNAfromProtQTLs.py DHRS7B ENSG00000109016 17:21036822nom_names=c("gene", "snp", "dist", "pval", "slope")
DHRS7B_RNA=read.table("../data/pQTL_otherphen/DHRS7B_17:21036822_RNAres.txt", col.names = nom_names, stringsAsFactors = F) %>% select(snp, slope) %>% filter(!duplicated(snp))
DHRS7B_Prot=read.table("../data/pQTL_otherphen/DHRS7B_17:21036822_Protres.txt", col.names = nom_names, stringsAsFactors = F)%>% select(snp, slope) %>% filter(!duplicated(snp))
#FILTER snps in the list from Rna and prot
DHRS7B_apaTot=read.table("../data/pQTL_otherphen/DHRS7B_17:21036822_Totalres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp))
DHRS7B_apaNuc=read.table("../data/pQTL_otherphen/DHRS7B_17:21036822_Nuclearres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp)) Start with RNA and protein because they are the most simple
DHRS7B_ProtRNA=DHRS7B_RNA %>% left_join(DHRS7B_Prot, by="snp")
colnames(DHRS7B_ProtRNA)=c("snp", "RNA", "Protein")Model:
DHRS7B_lmProtRNA=lm(Protein ~ RNA, data=DHRS7B_ProtRNA)
DHRS7B_lmProtRNA_sum=glance(DHRS7B_lmProtRNA)For APA I need to get a data frame that has the snps by the effect sizes for each peak.I basically want to spread this df.
DHRS7B_apaTot_s=spread(DHRS7B_apaTot, "peak", "slope",drop = F)DHRS7B_apaNuc_s=spread(DHRS7B_apaNuc, "peak", "slope",drop = F)I can look at the protein ~ apaTotal.
DHRS7B_apaTotProt=DHRS7B_apaTot_s %>% left_join(DHRS7B_Prot, by="snp")
colnames(DHRS7B_apaTotProt)=c(names(DHRS7B_apaTot_s),"Protein")DHRS7B_lmProtAPA=lm(Protein ~ peak132690+peak132692+peak132693+peak132694+peak132720+peak132721+peak132722+peak132723+peak132724+peak132725+peak132728+peak132729+peak132730+peak132732+peak132733+peak132734+peak132735+peak132738+peak132739+peak132740+peak132741+peak132742+peak132744+peak132745 +peak132746+peak132747+peak132748+peak132749+peak132750+peak132751+peak132753+peak132754+ peak132755+peak132756+peak132757+peak132760+peak132761+peak132762+peak132763+peak132764+peak132766+peak132774+peak132775+peak132777+peak132778+peak132780, data=DHRS7B_apaTotProt)
DHRS7B_lmProtAPA_sum=glance(DHRS7B_lmProtAPA)Try with all:
DHRS7B_apaTotProtRNA=DHRS7B_apaTot_s %>% inner_join(DHRS7B_ProtRNA, by="snp")
colnames(DHRS7B_apaTotProtRNA)=c(names(DHRS7B_apaTot_s),"RNA", "protein")DHRS7B_lmProtAPARNA=lm(protein ~ peak132690+peak132692+peak132693+peak132694+peak132720+peak132721+peak132722+peak132723+peak132724+peak132725+peak132728+peak132729+peak132730+peak132732+peak132733+peak132734+peak132735+peak132738+peak132739+peak132740+peak132741+peak132742+peak132744+peak132745 +peak132746+peak132747+peak132748+peak132749+peak132750+peak132751+peak132753+peak132754+ peak132755+peak132756+peak132757+peak132760+peak132761+peak132762+peak132763+peak132764+peak132766+peak132774+peak132775+peak132777+peak132778+peak132780, data=DHRS7B_apaTotProtRNA)
DHRS7B_lmProtAPARNA_sum=glance(DHRS7B_lmProtAPARNA)
adjR2_DHRS7B=cbind("DHRS7B",DHRS7B_lmProtRNA_sum[1,2], DHRS7B_lmProtAPA_sum[1,2], DHRS7B_lmProtAPARNA_sum[1,2])
colnames(adjR2_DHRS7B)=c("Gene", "RNA", "APA", "RNAandAPA")Example MMAB
ENSG00000139428 MMAB 12:109997847
MMAB_APAandRNAfromProtQTLs.sh
#!/bin/bash
#SBATCH --job-name=MMAB_APAandRNAfromProtQTLs
#SBATCH --account=pi-yangili1
#SBATCH --time=24:00:00
#SBATCH --output=MMAB_APAandRNAfromProtQTLs.out
#SBATCH --error=MMAB_APAandRNAfromProtQTLs.err
#SBATCH --partition=broadwl
#SBATCH --mem=12G
#SBATCH --mail-type=END
module load python
python APAandRNAfromProtQTLs.py MMAB ENSG00000139428 12:109997847 nom_names=c("gene", "snp", "dist", "pval", "slope")
MMAB_RNA=read.table("../data/pQTL_otherphen/MMAB_12:109997847_RNAres.txt", col.names = nom_names, stringsAsFactors = F) %>% select(snp, slope) %>% filter(!duplicated(snp))
MMAB_Prot=read.table("../data/pQTL_otherphen/MMAB_12:109997847_Protres.txt", col.names = nom_names, stringsAsFactors = F)%>% select(snp, slope) %>% filter(!duplicated(snp))
#FILTER snps in the list from Rna and prot
MMAB_apaTot=read.table("../data/pQTL_otherphen/MMAB_12:109997847_Totalres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp))
MMAB_apaNuc=read.table("../data/pQTL_otherphen/MMAB_12:109997847_Nuclearres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp)) Start with RNA and protein because they are the most simple
MMAB_ProtRNA=MMAB_RNA %>% left_join(MMAB_Prot, by="snp")
colnames(MMAB_ProtRNA)=c("snp", "RNA", "Protein")Model:
MMAB_lmProtRNA=lm(Protein ~ RNA, data=MMAB_ProtRNA)
MMAB_lmProtRNA_sum=glance(MMAB_lmProtRNA)For APA I need to get a data frame that has the snps by the effect sizes for each peak.I basically want to spread this df.
MMAB_apaTot_s=spread(MMAB_apaTot, "peak", "slope",drop = F)MMAB_apaNuc_s=spread(MMAB_apaNuc, "peak", "slope",drop = F)I can look at the protein ~ apaTotal.
MMAB_apaTotProt=MMAB_apaTot_s %>% left_join(MMAB_Prot, by="snp")
colnames(MMAB_apaTotProt)=c(names(MMAB_apaTot_s),"Protein")MMAB_lmProtAPA=lm(Protein ~ peak78754+peak78755+peak78756+peak78757+peak78758+peak78759+peak78760+peak78761+peak78762+peak78763+peak78764+peak78765+peak78766, data=MMAB_apaTotProt)
MMAB_lmProtAPA_sum=glance(MMAB_lmProtAPA)Try with all:
MMAB_apaTotProtRNA=MMAB_apaTot_s %>% inner_join(MMAB_ProtRNA, by="snp")
colnames(MMAB_apaTotProtRNA)=c(names(MMAB_apaTot_s),"RNA", "protein")MMAB_lmProtAPARNA=lm(protein ~RNA+ peak78754+peak78755+peak78756+peak78757+peak78758+peak78759+peak78760+peak78761+peak78762+peak78763+peak78764+peak78765+peak78766, data=MMAB_apaTotProtRNA)
MMAB_lmProtAPARNA_sum=glance(MMAB_lmProtAPARNA)
adjR2_MMAB=cbind("MMAB",MMAB_lmProtRNA_sum[1,2], MMAB_lmProtAPA_sum[1,2], MMAB_lmProtAPARNA_sum[1,2])
colnames(adjR2_MMAB)=c("Gene", "RNA", "APA", "RNAandAPA")Example FYTTD1
FYTTD1 ENSG00000122068 3:197035232
FYTTD1_APAandRNAfromProtQTLs.sh
#!/bin/bash
#SBATCH --job-name=FYTTD1_APAandRNAfromProtQTLs
#SBATCH --account=pi-yangili1
#SBATCH --time=24:00:00
#SBATCH --output=FYTTD1_APAandRNAfromProtQTLs.out
#SBATCH --error=FYTTD1_APAandRNAfromProtQTLs.err
#SBATCH --partition=broadwl
#SBATCH --mem=12G
#SBATCH --mail-type=END
module load python
python APAandRNAfromProtQTLs.py FYTTD1 ENSG00000122068 3:197035232 nom_names=c("gene", "snp", "dist", "pval", "slope")
FYTTD1_RNA=read.table("../data/pQTL_otherphen/FYTTD1_3:197035232_RNAres.txt", col.names = nom_names, stringsAsFactors = F) %>% select(snp, slope) %>% filter(!duplicated(snp))
FYTTD1_Prot=read.table("../data/pQTL_otherphen/FYTTD1_3:197035232_Protres.txt", col.names = nom_names, stringsAsFactors = F)%>% select(snp, slope) %>% filter(!duplicated(snp))
#FILTER snps in the list from Rna and prot
FYTTD1_apaTot=read.table("../data/pQTL_otherphen/FYTTD1_3:197035232_Totalres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp))
FYTTD1_apaNuc=read.table("../data/pQTL_otherphen/FYTTD1_3:197035232_Nuclearres.txt", col.names = nom_names, stringsAsFactors = F) %>% separate(gene, sep = ":", into=c("chr", "start", "end", "id")) %>% separate(id, sep = "_", into=c("gene", "strand", "peak")) %>% select(snp,peak, slope) %>% group_by(peak) %>% filter(!duplicated(snp)) Start with RNA and protein because they are the most simple
FYTTD1_ProtRNA=FYTTD1_RNA %>% left_join(FYTTD1_Prot, by="snp")
colnames(FYTTD1_ProtRNA)=c("snp", "RNA", "Protein")Model:
FYTTD1_lmProtRNA=lm(Protein ~ RNA, data=FYTTD1_ProtRNA)
FYTTD1_lmProtRNA_sum=glance(FYTTD1_lmProtRNA)For APA I need to get a data frame that has the snps by the effect sizes for each peak.I basically want to spread this df.
FYTTD1_apaTot_s=spread(FYTTD1_apaTot, "peak", "slope",drop = F)FYTTD1_apaNuc_s=spread(FYTTD1_apaNuc, "peak", "slope",drop = F)I can look at the protein ~ apaTotal.
FYTTD1_apaTotProt=FYTTD1_apaTot_s %>% left_join(FYTTD1_Prot, by="snp")
colnames(FYTTD1_apaTotProt)=c(names(FYTTD1_apaTot_s),"Protein")FYTTD1_lmProtAPA=lm(Protein ~ peak234016 +peak234082 +peak234088+peak234089+peak234090+peak234092+peak234093+peak234094+peak234095+peak234099+peak234101+peak234103+peak234104+peak234105+peak234106+peak234107+peak234108+peak234111+peak234113+peak234114+peak234117+peak234119+peak234120+peak234122+peak234123+peak234124+peak234125+peak234131+peak234135, data=FYTTD1_apaTotProt)
FYTTD1_lmProtAPA_sum=glance(FYTTD1_lmProtAPA)FYTTD1_apaTotProtRNA=FYTTD1_apaTot_s %>% inner_join(FYTTD1_ProtRNA, by="snp")
colnames(FYTTD1_apaTotProtRNA)=c(names(FYTTD1_apaTot_s),"RNA", "protein")FYTTD1_lmProtAPARNA=lm(protein ~RNA+ peak234016 +peak234082 +peak234088+peak234089+peak234090+peak234092+peak234093+peak234094+peak234095+peak234099+peak234101+peak234103+peak234104+peak234105+peak234106+peak234107+peak234108+peak234111+peak234113+peak234114+peak234117+peak234119+peak234120+peak234122+peak234123+peak234124+peak234125+peak234131+peak234135, data=FYTTD1_apaTotProtRNA)
FYTTD1_lmProtAPARNA_sum=glance(FYTTD1_lmProtAPARNA)
adjR2_FYTTD1=cbind("FYTTD1",FYTTD1_lmProtRNA_sum[1,2], FYTTD1_lmProtAPA_sum[1,2], FYTTD1_lmProtAPARNA_sum[1,2])
colnames(adjR2_FYTTD1)=c("Gene", "RNA", "APA", "RNAandAPA")Analysis
In each of these examples the total APA explains more of the protein variability. I am going to plot the adjR2 for each gene and model
adjR2=rbind(adjR2_CCDC51,adjR2_DTD1,adjR2_DHRS7B, adjR2_MMAB, adjR2_FYTTD1)
adjR2_melt=melt(adjR2, id.vars="Gene")
colnames(adjR2_melt)=c("Gene", "Model", "AdjR2")Plot this
ggplot(adjR2_melt, aes(x=Gene, y=AdjR2, by=Model, fill=Model)) + geom_bar(stat="identity", position = "dodge") + labs(title="Adjusted R2 for variation\n in protein effect sizes explained in each model",x="pQTL Gene") + scale_fill_brewer(palette="Paired")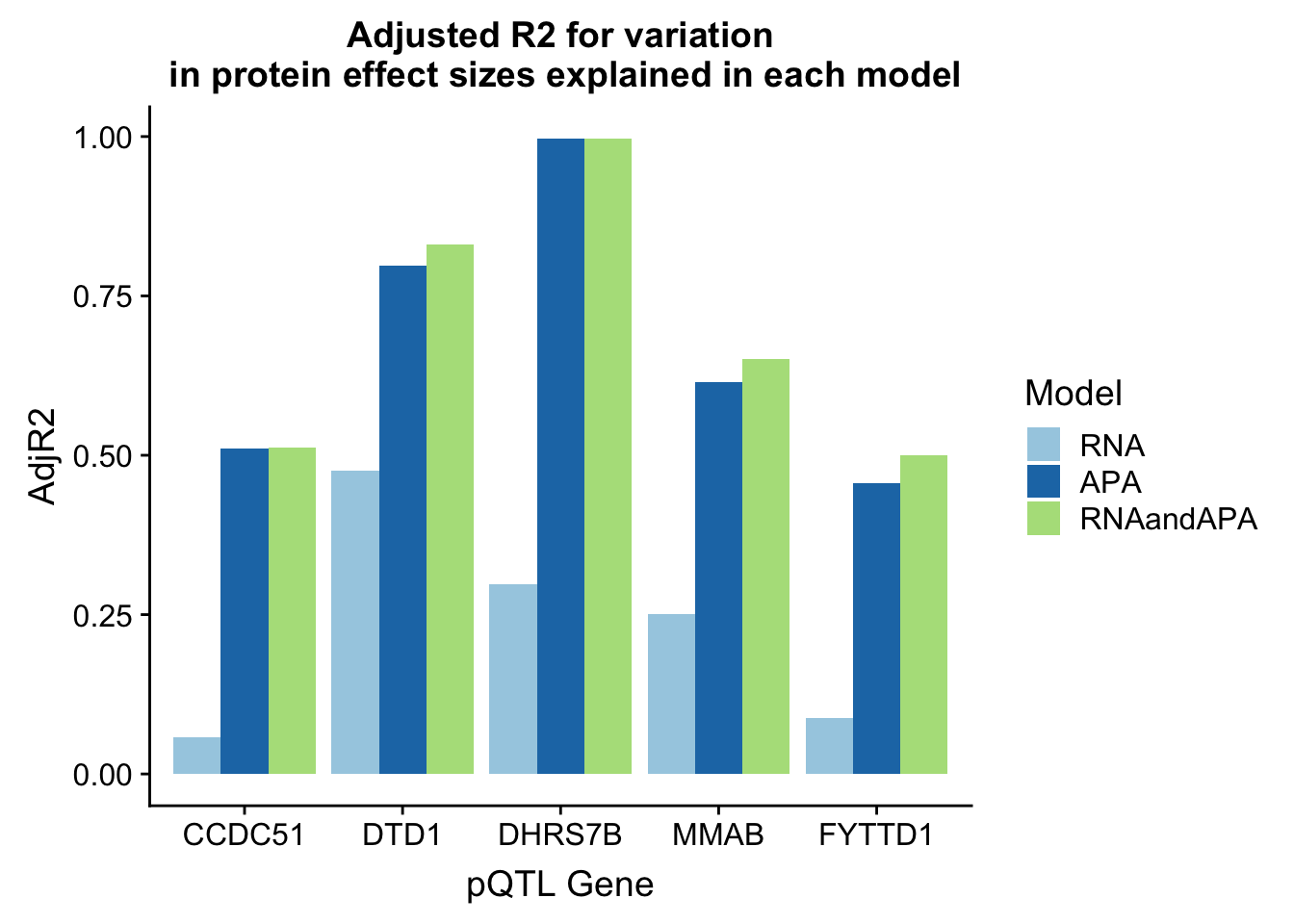
Session information
sessionInfo()R version 3.5.1 (2018-07-02)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS 10.14.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.5/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] bindrcpp_0.2.2 cowplot_0.9.3 broom_0.5.0 reshape2_1.4.3
[5] forcats_0.3.0 stringr_1.3.1 dplyr_0.7.6 purrr_0.2.5
[9] readr_1.1.1 tidyr_0.8.1 tibble_1.4.2 ggplot2_3.0.0
[13] tidyverse_1.2.1 workflowr_1.1.1
loaded via a namespace (and not attached):
[1] tidyselect_0.2.4 haven_1.1.2 lattice_0.20-35
[4] colorspace_1.3-2 htmltools_0.3.6 yaml_2.2.0
[7] rlang_0.2.2 R.oo_1.22.0 pillar_1.3.0
[10] glue_1.3.0 withr_2.1.2 R.utils_2.7.0
[13] RColorBrewer_1.1-2 modelr_0.1.2 readxl_1.1.0
[16] bindr_0.1.1 plyr_1.8.4 munsell_0.5.0
[19] gtable_0.2.0 cellranger_1.1.0 rvest_0.3.2
[22] R.methodsS3_1.7.1 evaluate_0.11 labeling_0.3
[25] knitr_1.20 Rcpp_0.12.19 scales_1.0.0
[28] backports_1.1.2 jsonlite_1.5 hms_0.4.2
[31] digest_0.6.17 stringi_1.2.4 grid_3.5.1
[34] rprojroot_1.3-2 cli_1.0.1 tools_3.5.1
[37] magrittr_1.5 lazyeval_0.2.1 crayon_1.3.4
[40] whisker_0.3-2 pkgconfig_2.0.2 xml2_1.2.0
[43] lubridate_1.7.4 assertthat_0.2.0 rmarkdown_1.10
[46] httr_1.3.1 rstudioapi_0.8 R6_2.3.0
[49] nlme_3.1-137 git2r_0.23.0 compiler_3.5.1
This reproducible R Markdown analysis was created with workflowr 1.1.1