Contents
close all,
chooseFigure = 6;
saveFigure = 0; plotFigure = 1; maxTrial = 10;
maxStart = 10;
computeCostBernoulli = @(X, P)(sum(sum(-X.*log(double(P + eps)))) ...
+ sum(sum(-(1 - X).*log(1 - double(P) + eps))) );
computeCostPoisson = @(X, R)(sum(sum(- X .* log(R + eps) + R)));
computeCostStochastic = @(X, P)...
(- sum(sum(X .* log(P + eps),2)));
Multinomial data
rng default, close all,
MARKERSIZE = 4; FONTSIZE = 6;
if plotFigure; hMain = figure('papersize',[8.5 11],'paperposition',[0 0 8 2]); end;
setenv('DYLD_LIBRARY_PATH', '/usr/local/bin/');
d = 3;
K = 5;
n = 500;
colorList = summer(maxTrial);
gammaParam = 0.5;
if d ~= 3
fprintf('Only valid for d = 3.\n')
end
matFeatLat = convexCircle(K);
matLatSam = gamrnd(gammaParam, 1, K, n); matLatSam = bsxfun(@rdivide, matLatSam, sum(matLatSam));
matFeatSam = matFeatLat * matLatSam;
nFeatSam = mnrnd(ceil(1000*rand(n,1) + 1000), matFeatSam')';
options = generate_options();
options.maxIter = 1000;
options.verbose = false;
options.eps = 10^-6;
options.display = 0;
matFeatSam = bsxfun(@rdivide, nFeatSam, sum(nFeatSam));
l1List = zeros(maxTrial, 2); nllList = zeros(maxTrial, 2);
for countTrial = 1:maxTrial
matSamLatEM = cell(maxStart,1);
matLatSamEM = cell(maxStart,1);
objEM = cell(maxStart,1);
for count = 1:maxStart
[matSamLatEM{count}, matLatSamEM{count}, objEM{count}] = paa_stochastic(nFeatSam, K, options);
end
[~, ind] = min(cellfun(@(x)(x(end)), objEM));
matSamLatEM = matSamLatEM{ind};
matLatSamEM = matLatSamEM{ind};
if plotFigure
figure(hMain)
subplot(131),
if countTrial == 1
set(gca,'fontsize',FONTSIZE), box on
hold on
plot(matFeatSam(1,:),matFeatSam(2,:),'o','markersize',MARKERSIZE,'markerfacecolor',[0.5 0.5 0.5],'markeredgecolor','b')
end
if countTrial == maxTrial
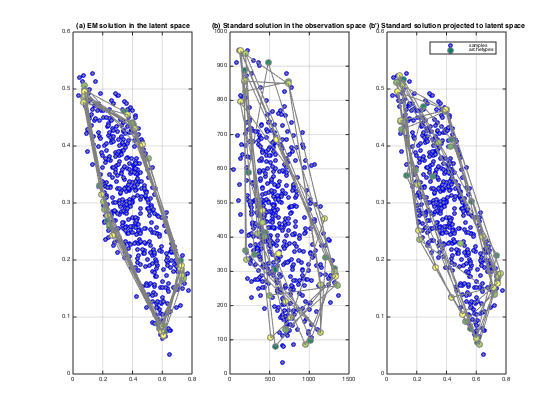
title('(a) EM solution in the latent space')
grid on
end
matFeatLatEM = matFeatSam * matSamLatEM;
plot(matFeatLatEM(1,:),matFeatLatEM(2,:),'ro','markersize',MARKERSIZE+2,'markerfacecolor',colorList(countTrial,:),'markeredgecolor',[0.5 0.5 0.5])
ind = convhull(matFeatLatEM(1:2,:)');
plot(matFeatLatEM(1,ind), matFeatLatEM(2,ind), '-', 'color', [0.5 0.5 0.5]);
end
if plotFigure
[matSamLatClassic, matLatSamClassic, objClassic] = classic_aa_plot(nFeatSam + rand(size(nFeatSam)), K);
else
[matSamLatClassic, matLatSamClassic, objClassic] = classic_aa(nFeatSam + rand(size(nFeatSam)), K);
end
if plotFigure
figure(hMain)
subplot(132),
if countTrial == 1
set(gca,'fontsize',FONTSIZE), box on
hold on
plot(nFeatSam(1,:),nFeatSam(2,:),'o','markersize',MARKERSIZE,'markerfacecolor',[0.5 0.5 0.5],'markeredgecolor','b')
end
if countTrial == maxTrial
title('(b) Standard solution in the observation space')
grid on
end
matFeatLatClassic = nFeatSam * matSamLatClassic;
plot(matFeatLatClassic(1,:),matFeatLatClassic(2,:),'ro','markersize',MARKERSIZE+2,'markerfacecolor',colorList(countTrial,:),'markeredgecolor',[0.5 0.5 0.5])
ind = convhull(matFeatLatClassic(1:2,:)');
plot(matFeatLatClassic(1,ind), matFeatLatClassic(2,ind), '-', 'color', [0.5 0.5 0.5]);
end
if plotFigure
figure(hMain)
subplot(133),
if countTrial == 1
set(gca,'fontsize',FONTSIZE), box on
hold on
plot(matFeatSam(1,:),matFeatSam(2,:),'o','markersize',MARKERSIZE,'markerfacecolor',[0.5 0.5 0.5],'markeredgecolor','b')
end
if countTrial == maxTrial
title('(b'') Standard solution projected to latent space')
grid on
end
matFeatLatClassic = matFeatSam * matSamLatClassic;
hold on, plot(matFeatLatClassic(1,:),matFeatLatClassic(2,:),'ro','markersize',MARKERSIZE+2,'markerfacecolor',colorList(countTrial,:),'markeredgecolor',[0.5 0.5 0.5])
ind = convhull(matFeatLatClassic(1:2,:)');
plot(matFeatLatClassic(1,ind), matFeatLatClassic(2,ind), '-', 'color', [0.5 0.5 0.5]);
end
if ~plotFigure
matFeatLatEM = matFeatSam * matSamLatEM;
matFeatLatClassic = matFeatSam * matSamLatClassic;
matFeatSam = matFeatLat * matLatSam;
nFeatSam = mnrnd(ceil(1000*rand(n,1) + 1000), matFeatSam')';
matFeatSam = bsxfun(@rdivide, nFeatSam, sum(nFeatSam));
clear matLatSamEM objEM
options.matFeatLat = matFeatLatEM;
for countEM = 1:maxStart
[~, matLatSamEM{countEM}, objEM{countEM}] = paa_stochastic(nFeatSam, K, options);
end
[~, ind] = min(cellfun(@(x)(x(end)), objEM));
matLatSamEM = matLatSamEM{ind};
nllList(countTrial, 1) = computeCostStochastic(nFeatSam, matFeatLatEM * matLatSamEM);
l1List(countTrial, 1) = norm(matFeatSam - matFeatLatEM * matLatSamEM);
options.matFeatLat = [];
matLatSamClassic = classic_aa_test(nFeatSam);
nllList(countTrial, 2) = computeCostStochastic(nFeatSam, matFeatLatClassic * matLatSamClassic);
l1List(countTrial, 2) = norm(matFeatSam - matFeatLatClassic * matLatSamClassic, 1);
end
end
if plotFigure
legend('samples','archetypes','location','NorthEast')
end
if saveFigure == 1
saveas(gcf,'compareProbVsStandard.eps','epsc')
!epstopdf compareProbVsStandard.eps
end
save compareProbVsDefault_stochastic nllList l1List
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1:
Loading required package: methods
Loading required package: modeltools
Loading required package: stats4
Loading required package: nnls
R.matlab v3.1.1 (2014-10-10) successfully loaded. See ?R.matlab for help.
Attaching package: 'R.matlab'
The following objects are masked from 'package:base':
getOption, isOpen
*** k=5, rep=1: