Simulation with Realistic Standard Error
Lei Sun
2017-05-27
Last updated: 2017-05-30
Code version: 9101229
library(ashr)
library(edgeR)
library(limma)
library(pROC)source("../code/gdash.R")Read in data
data = readRDS("../data/liver.rds")
ngene = 1e4
data = top_gene_selection(ngene, data)$dataGlobal Null
\[ g \equiv 0 \ . \]
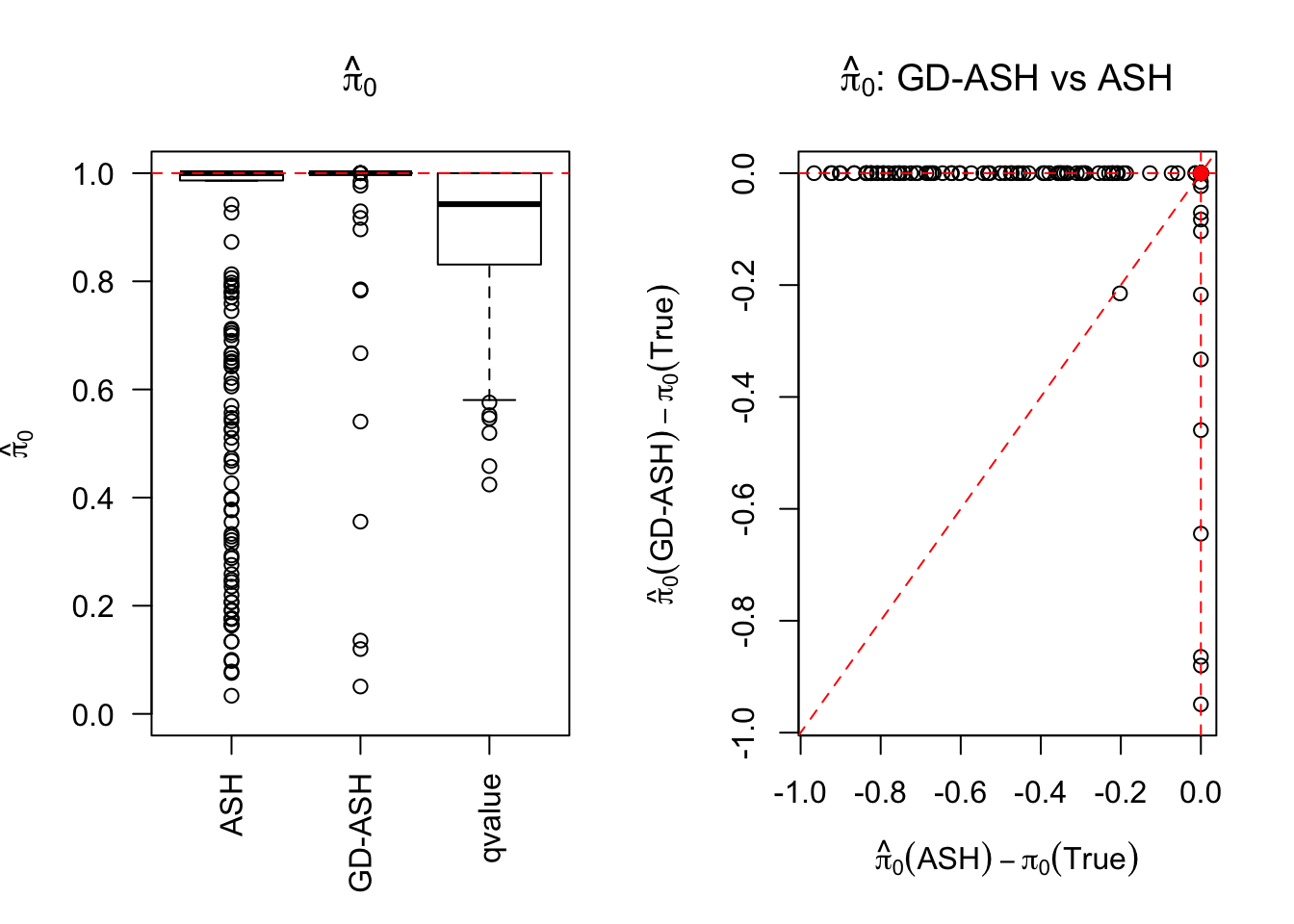
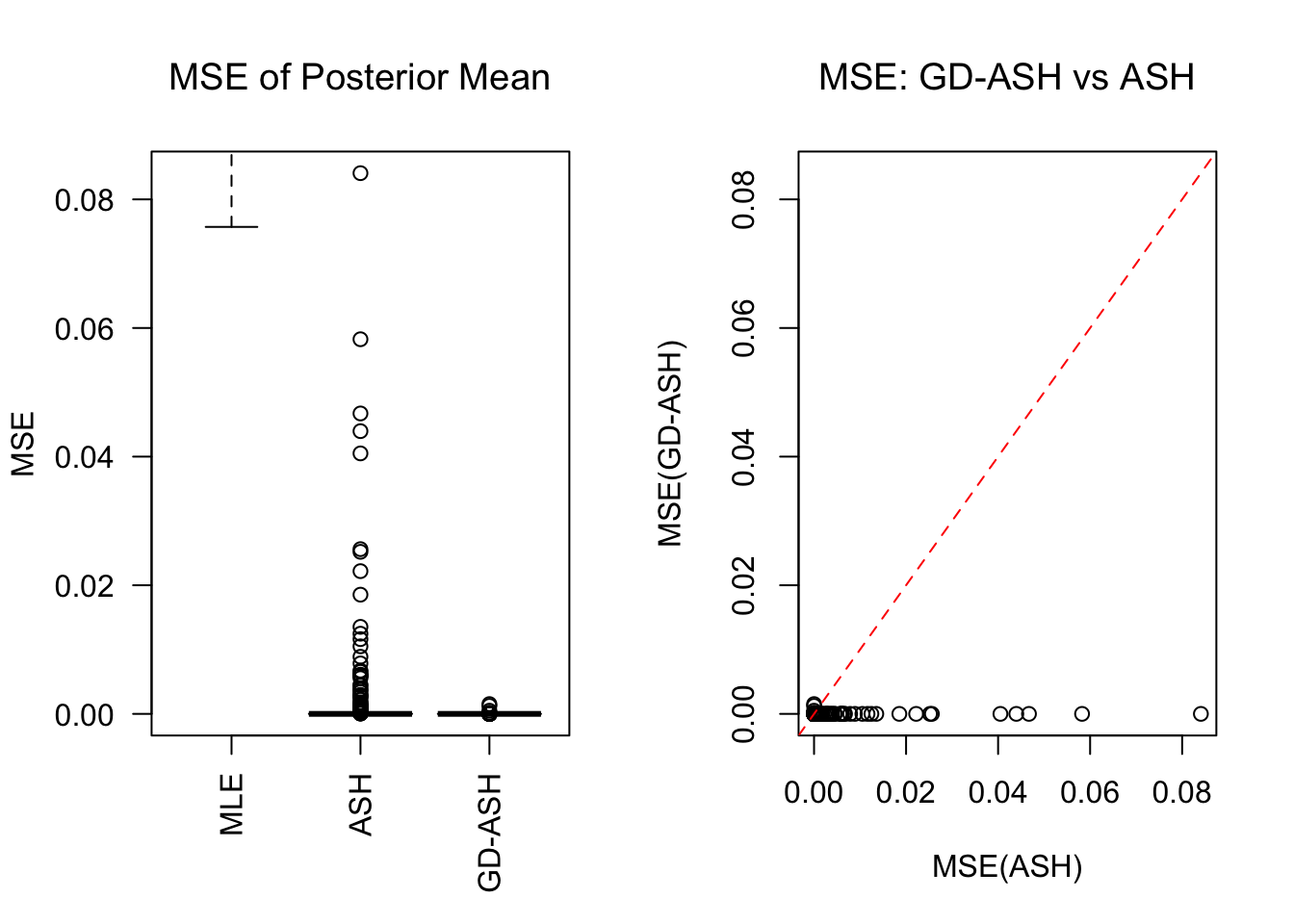
SNR = 0
\[ g = 0.6\delta_0 + 0.3N\left(0, \sigma^2\right) + 0.1N\left(0, \left(2.65\sigma\right)^2\right) \ . \]
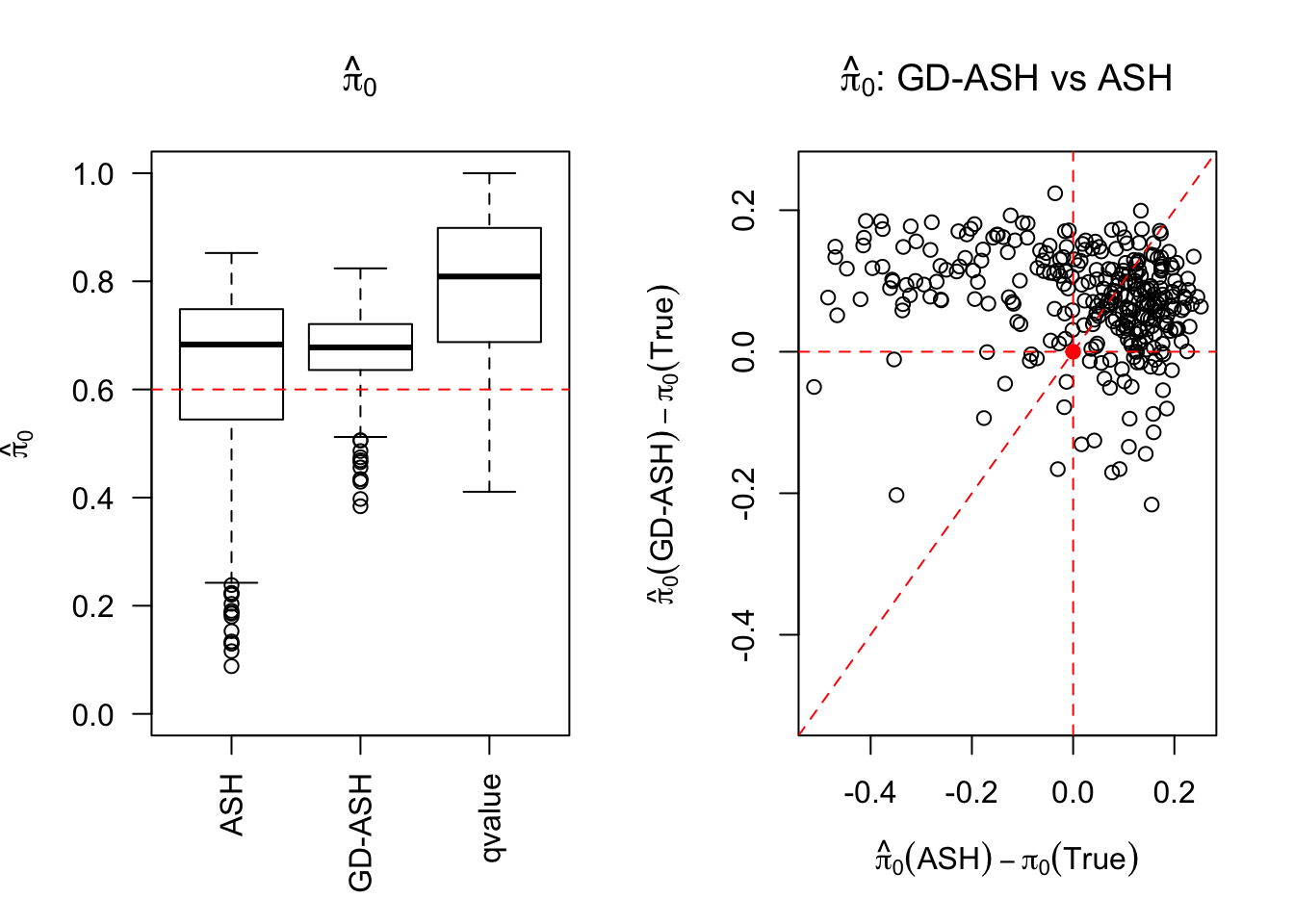
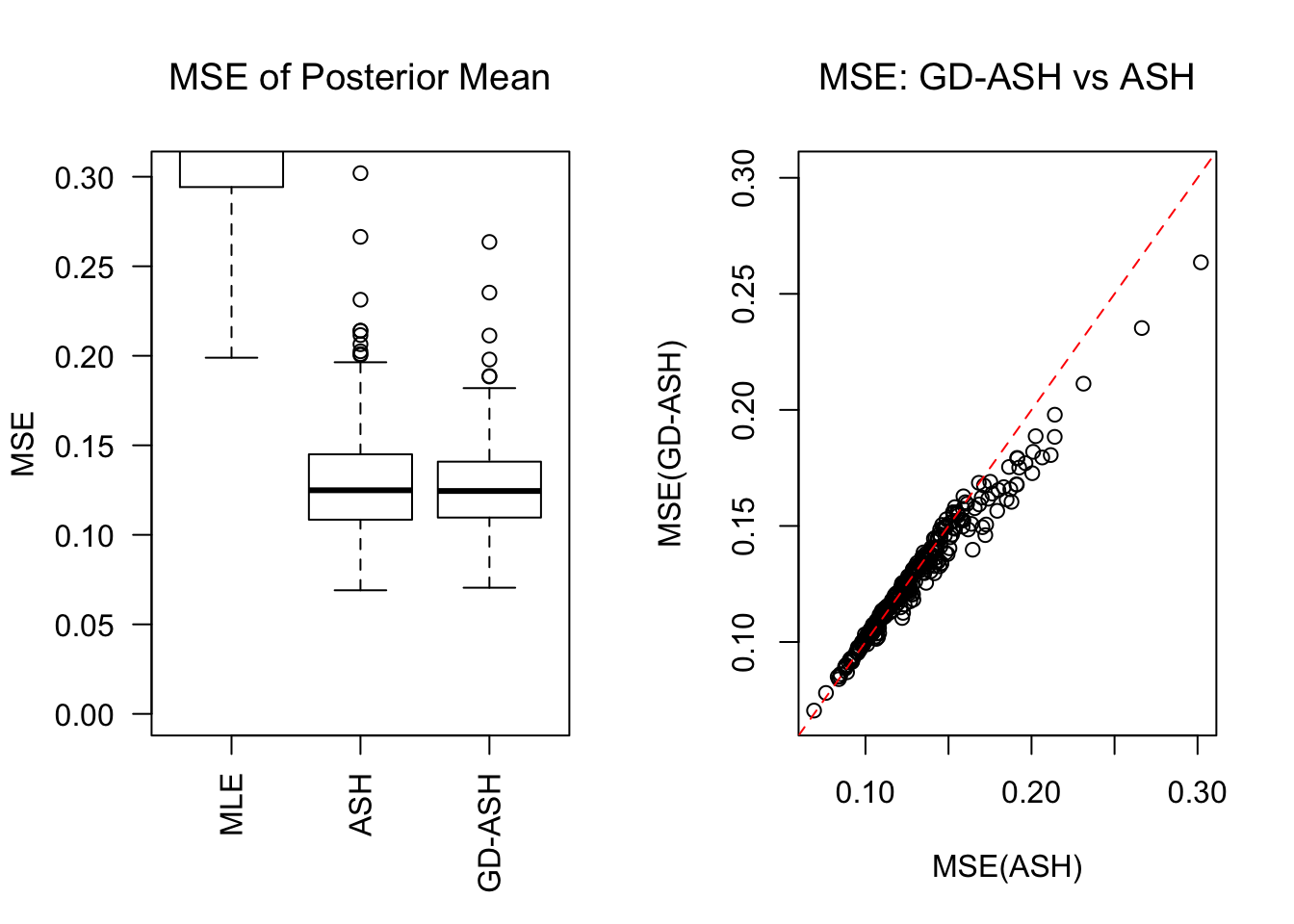
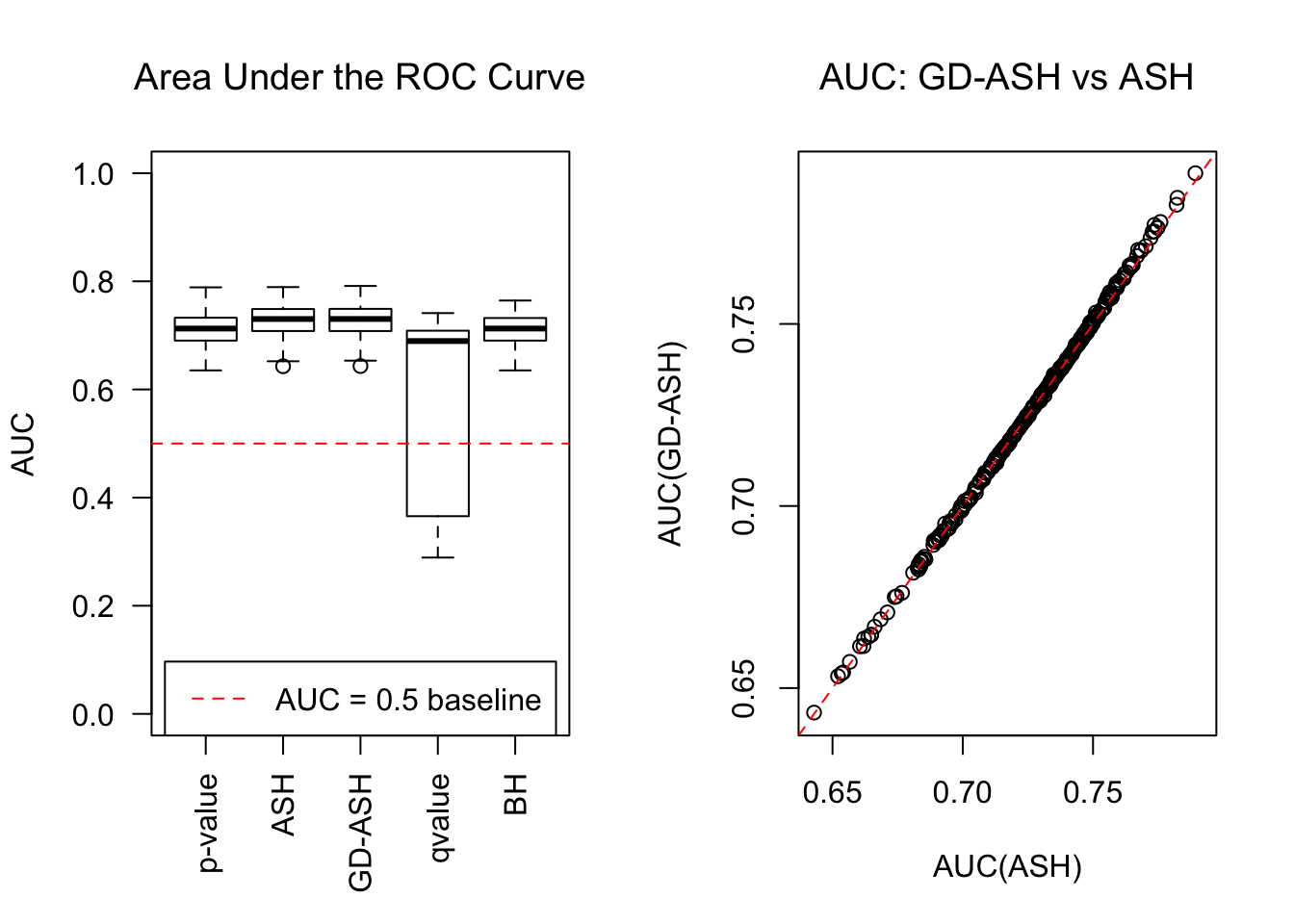
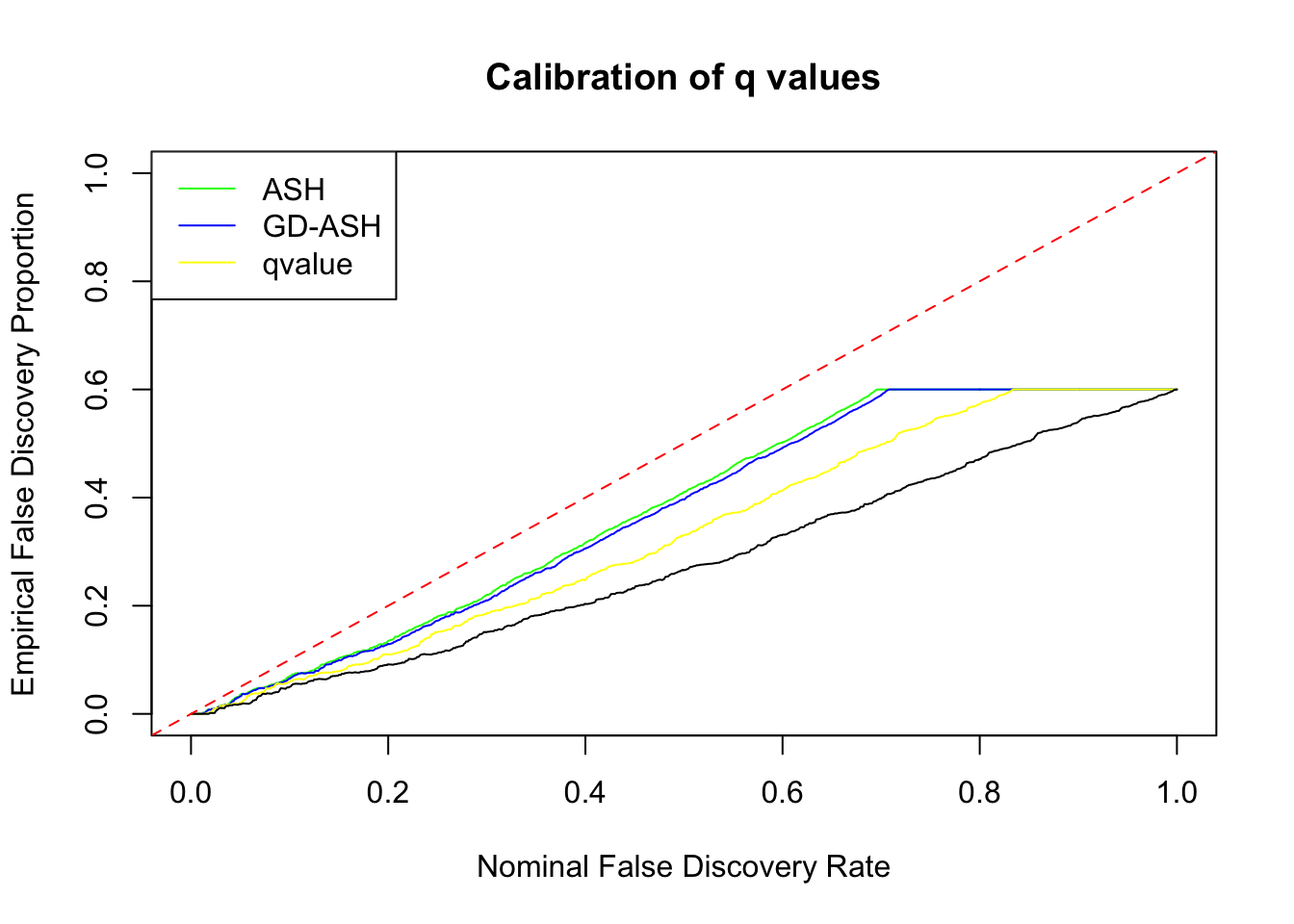
SNR = 2
\[ g = 0.6\delta_0 + 0.3N\left(0, \sigma^2\right) + 0.1N\left(0, \left(3.58\sigma\right)^2\right) \ . \]
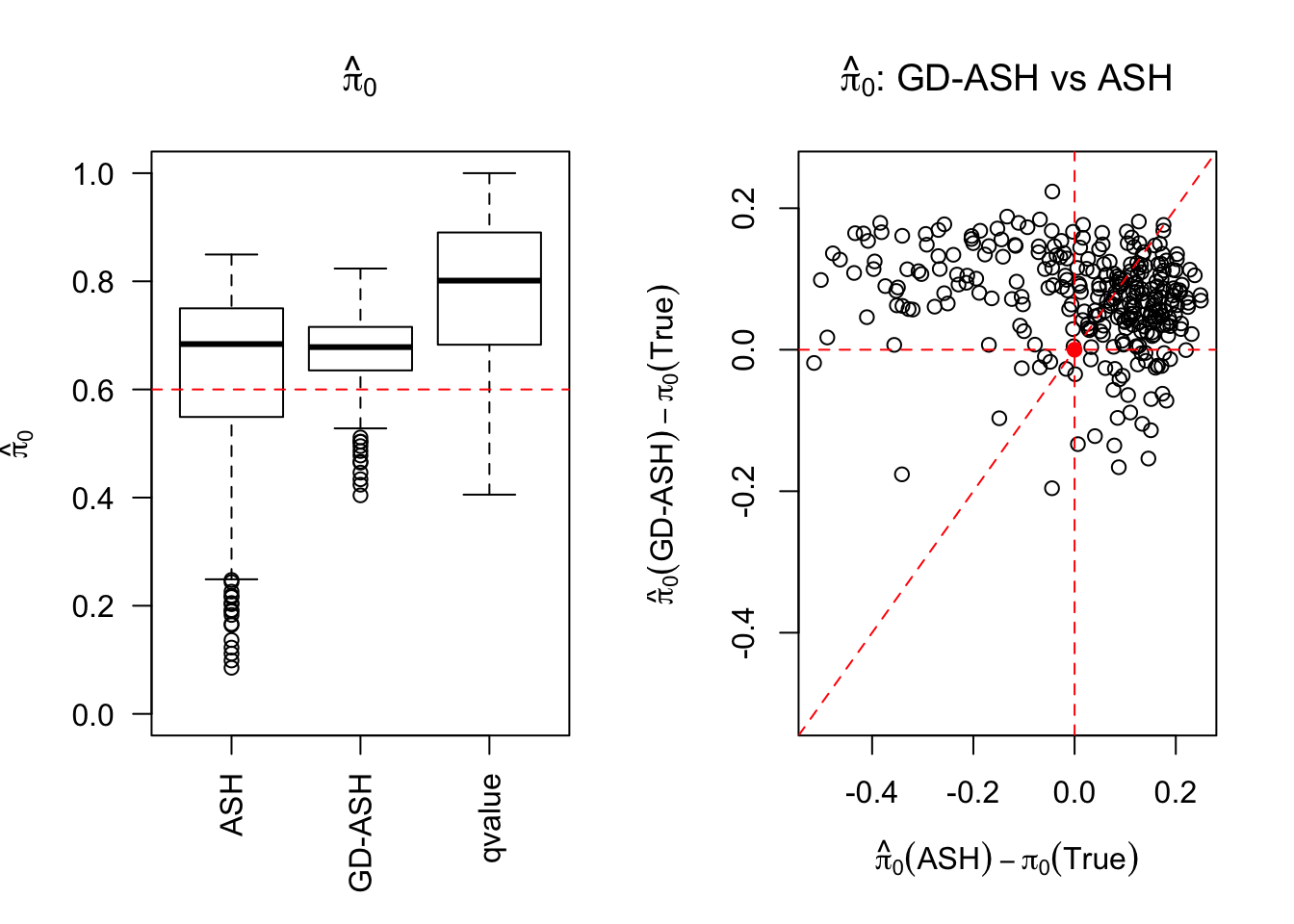
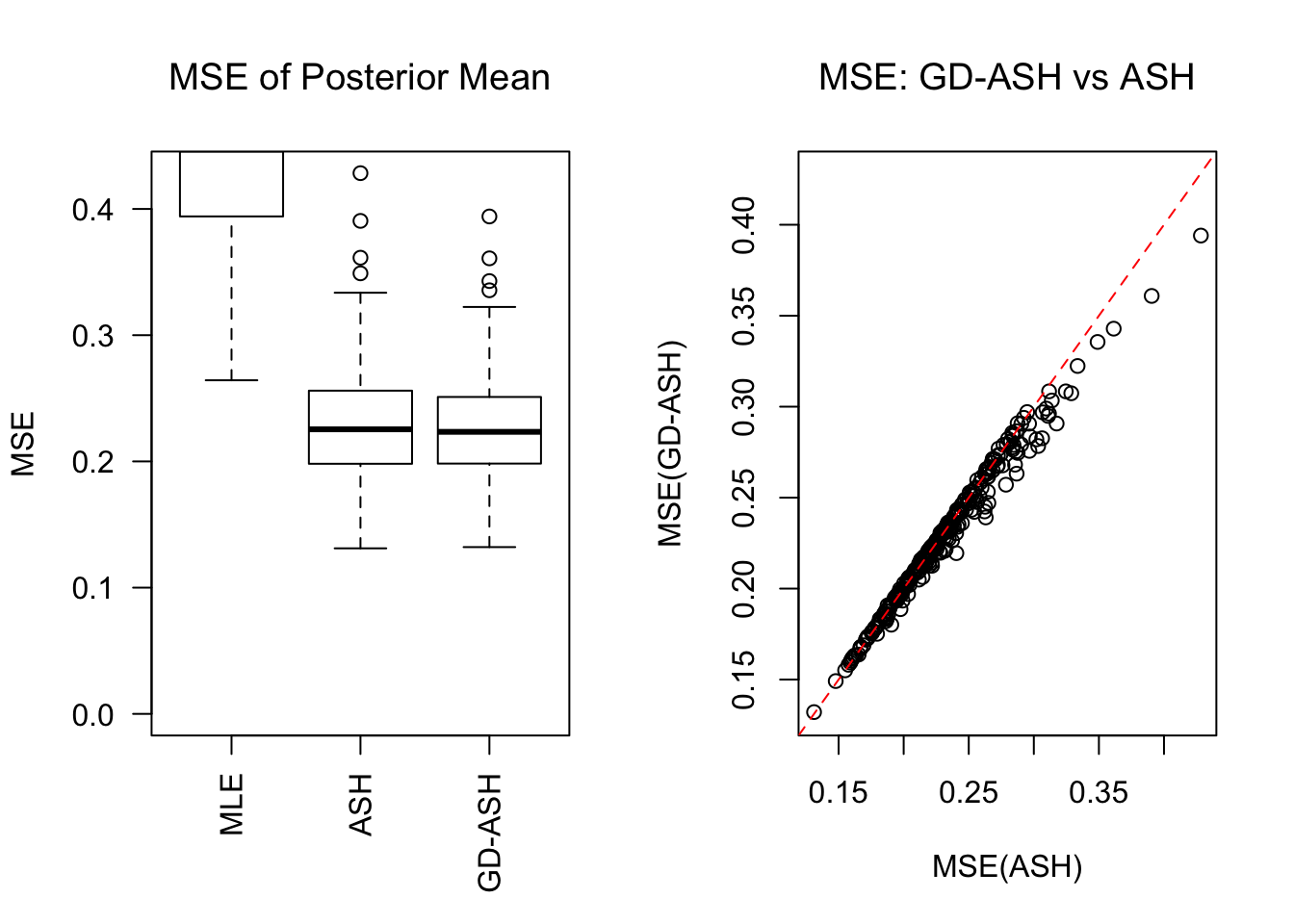
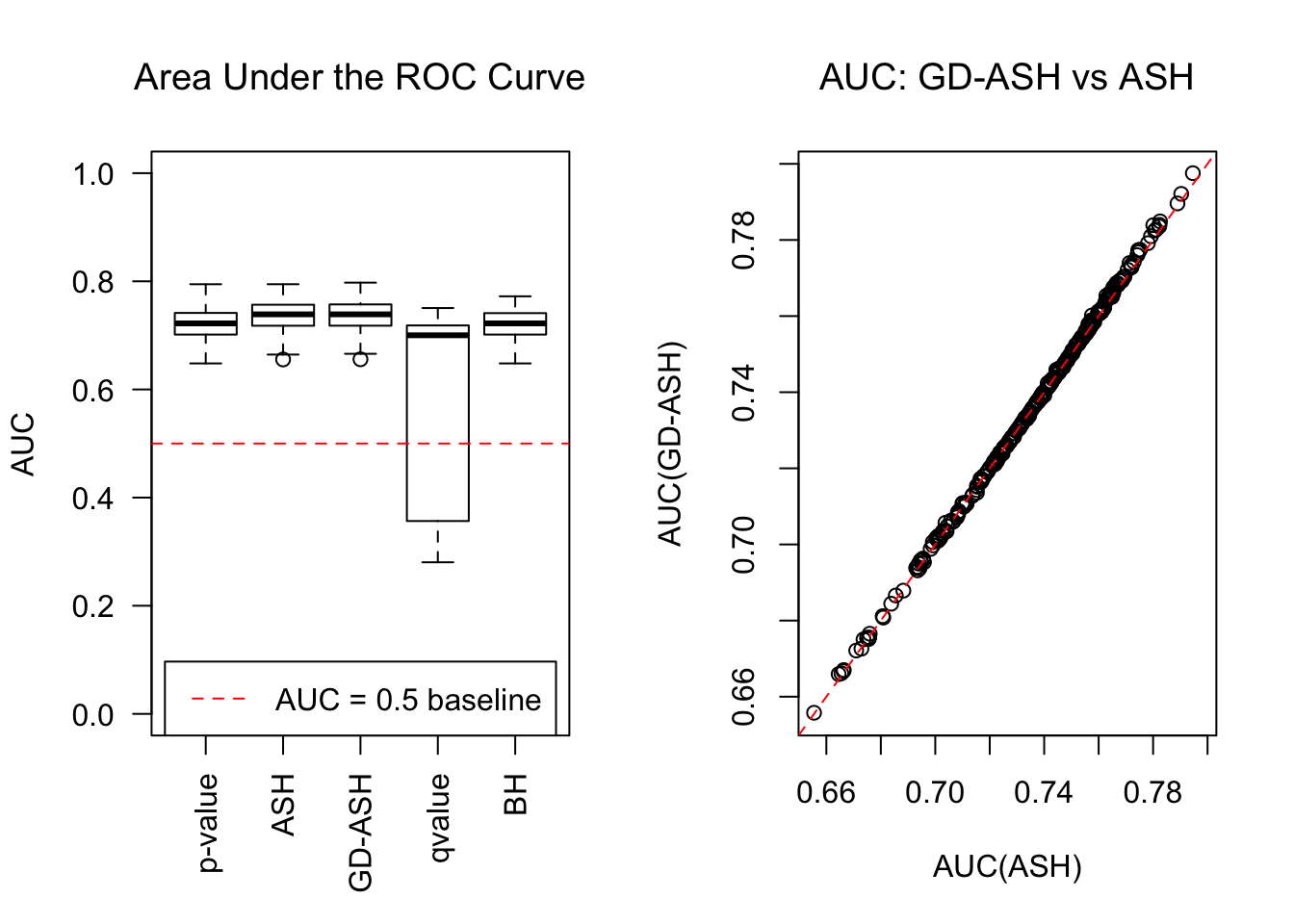
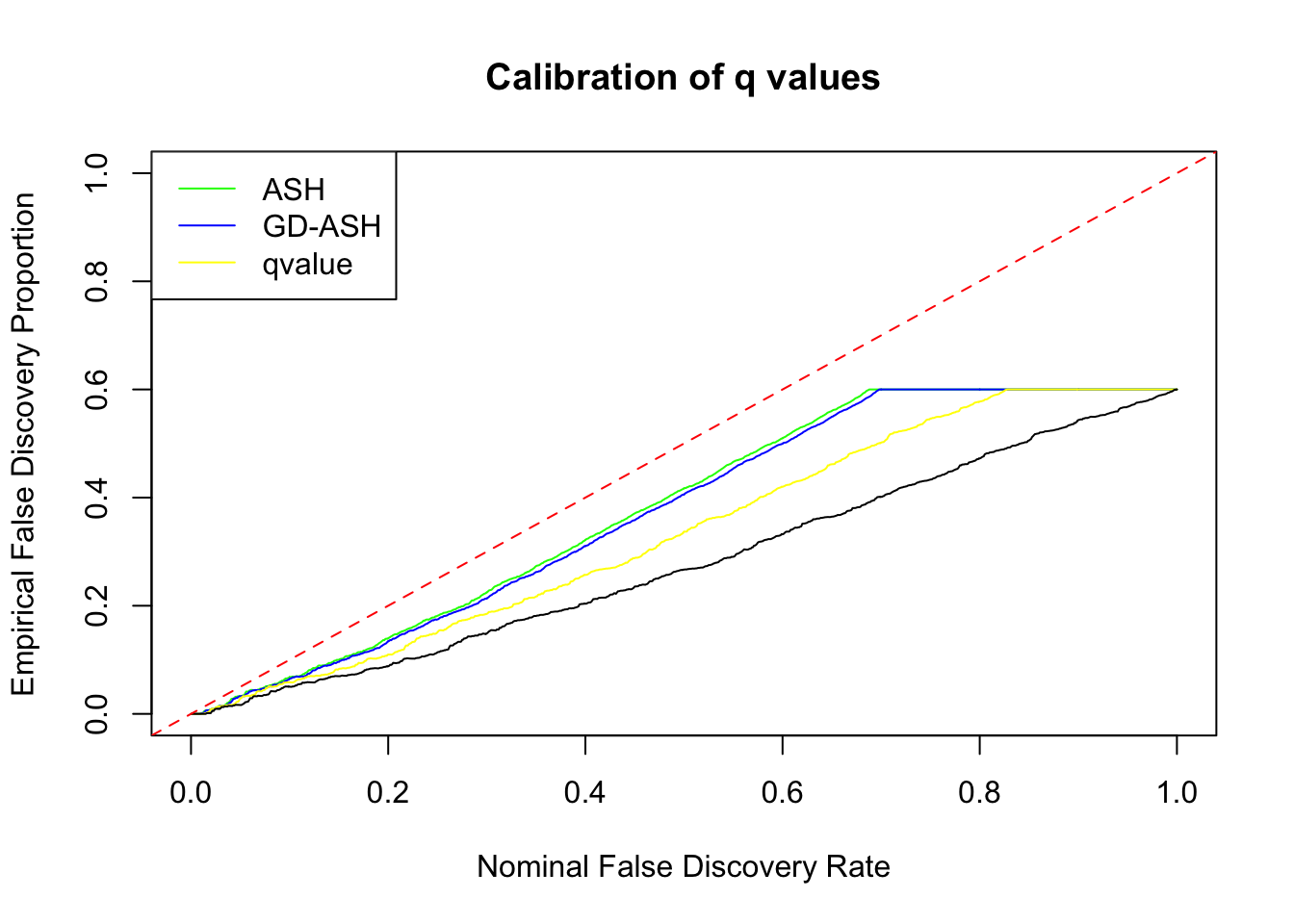
SNR = 5
\[ g = 0.6\delta_0 + 0.3N\left(0, \left(2\sigma\right)^2\right) + 0.1N\left(0, \left(4.43\sigma\right)^2\right) \ . \]
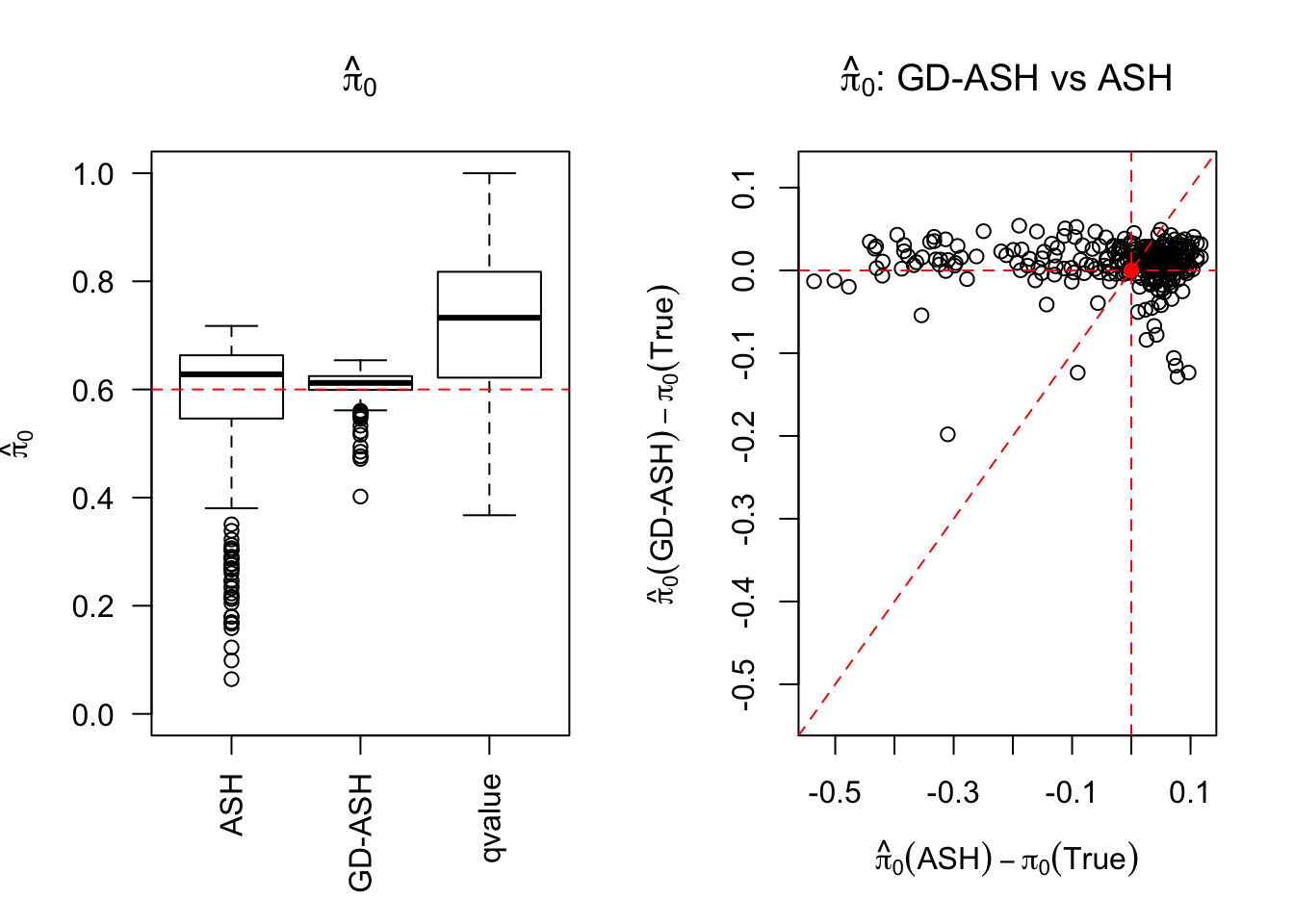
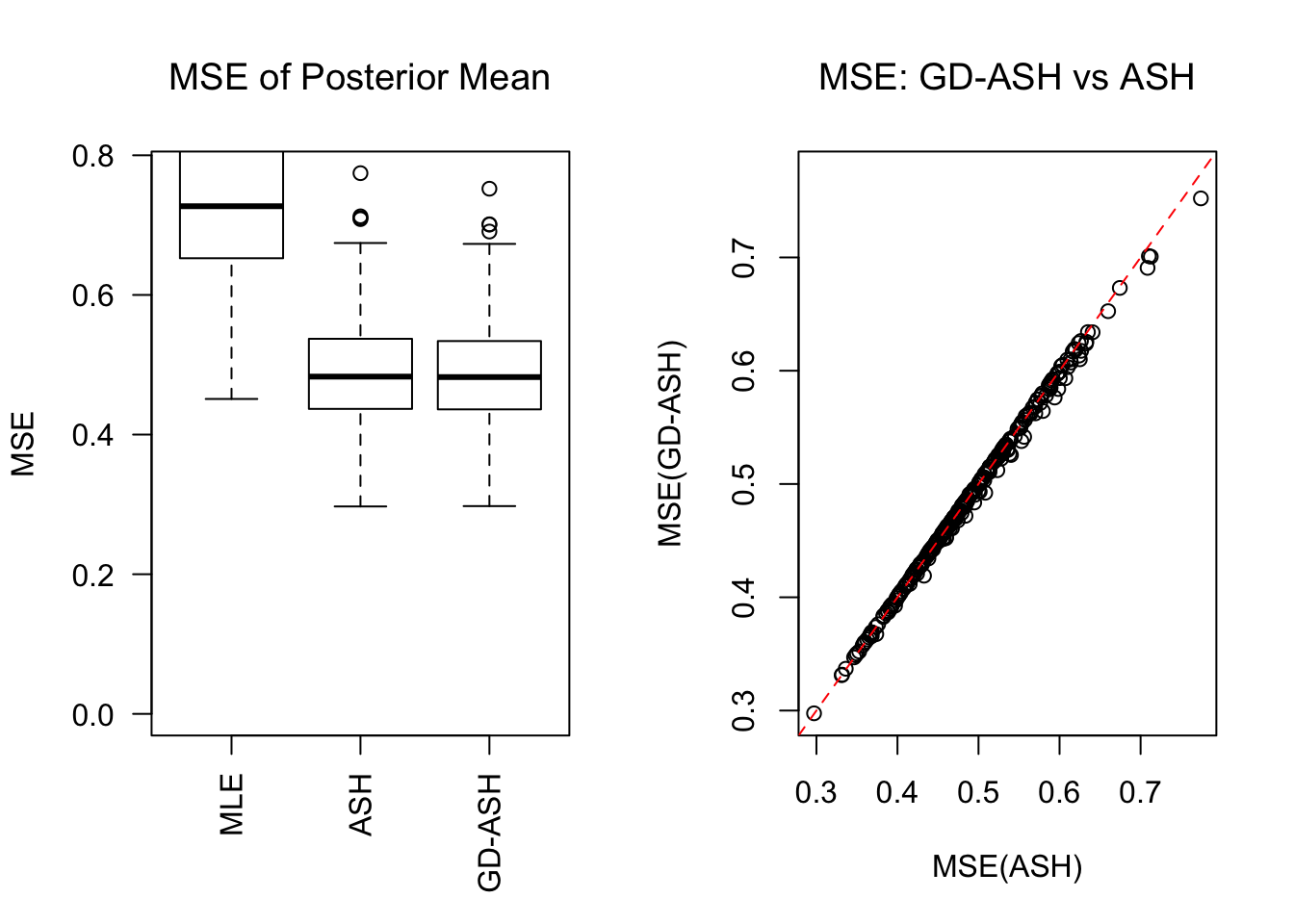
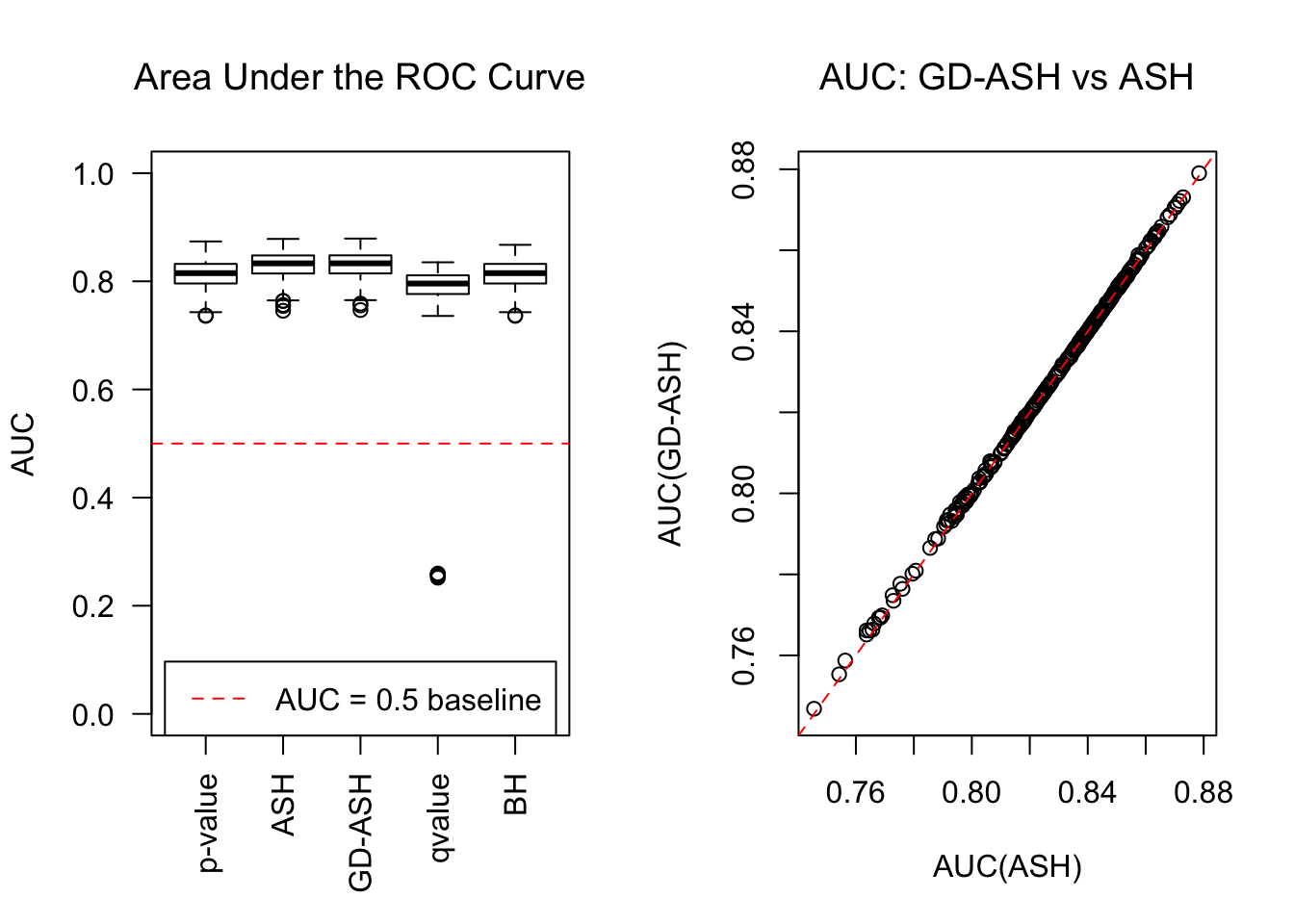
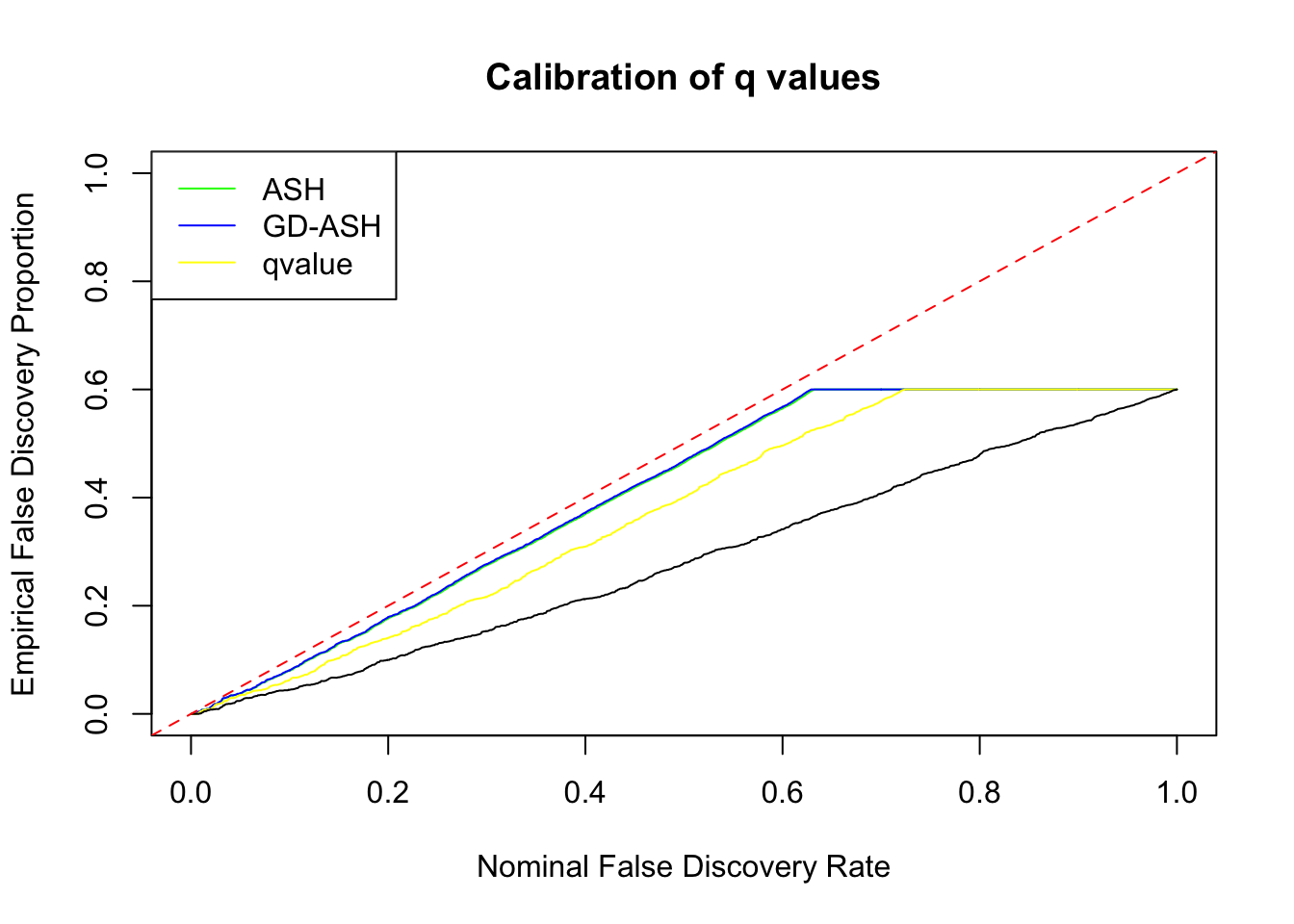
SNR = 10
\[ g = 0.6\delta_0 + 0.3N\left(0, \left(3\sigma\right)^2\right) + 0.1N\left(0, \left(8.54\sigma\right)^2\right) \ . \]
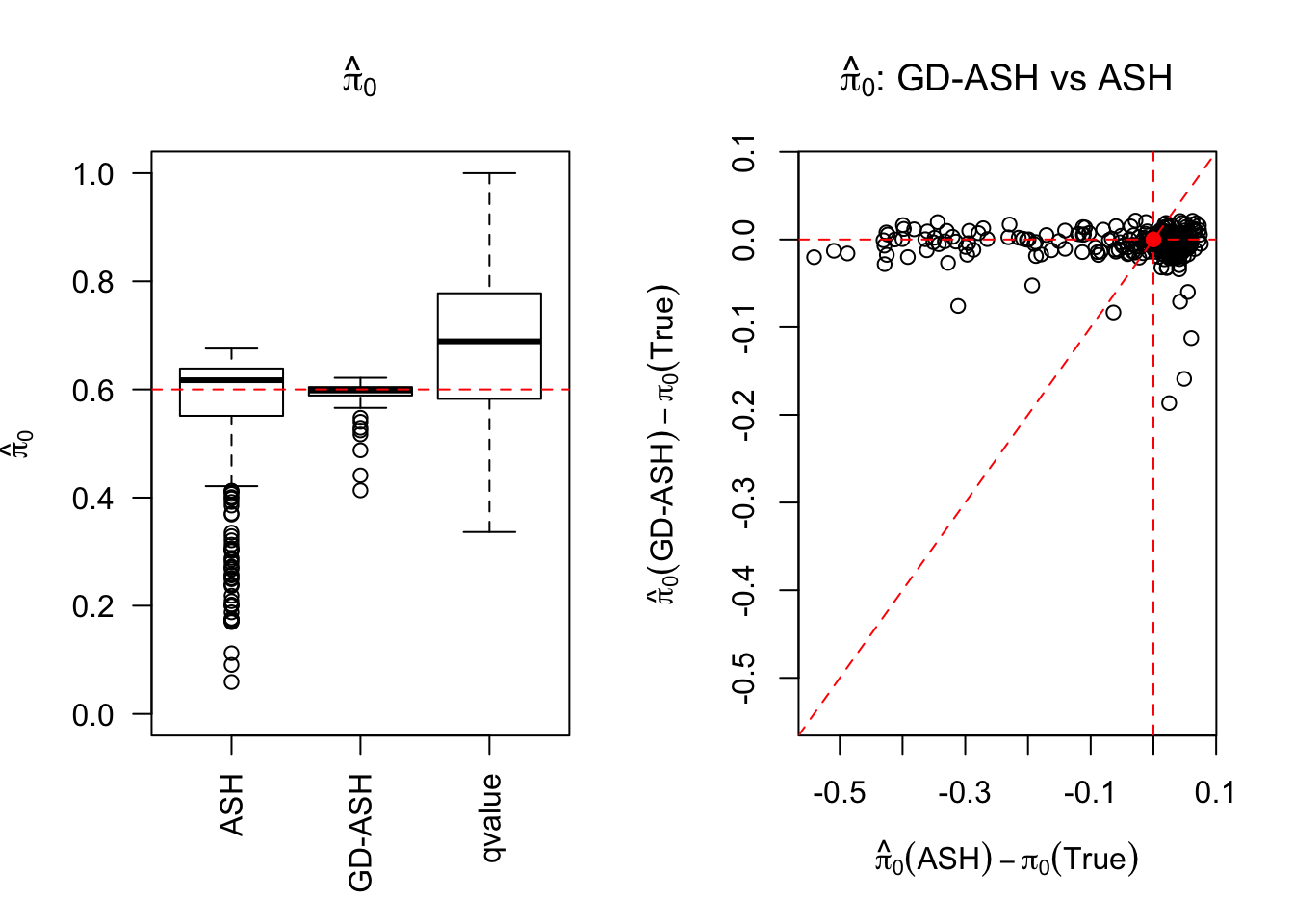
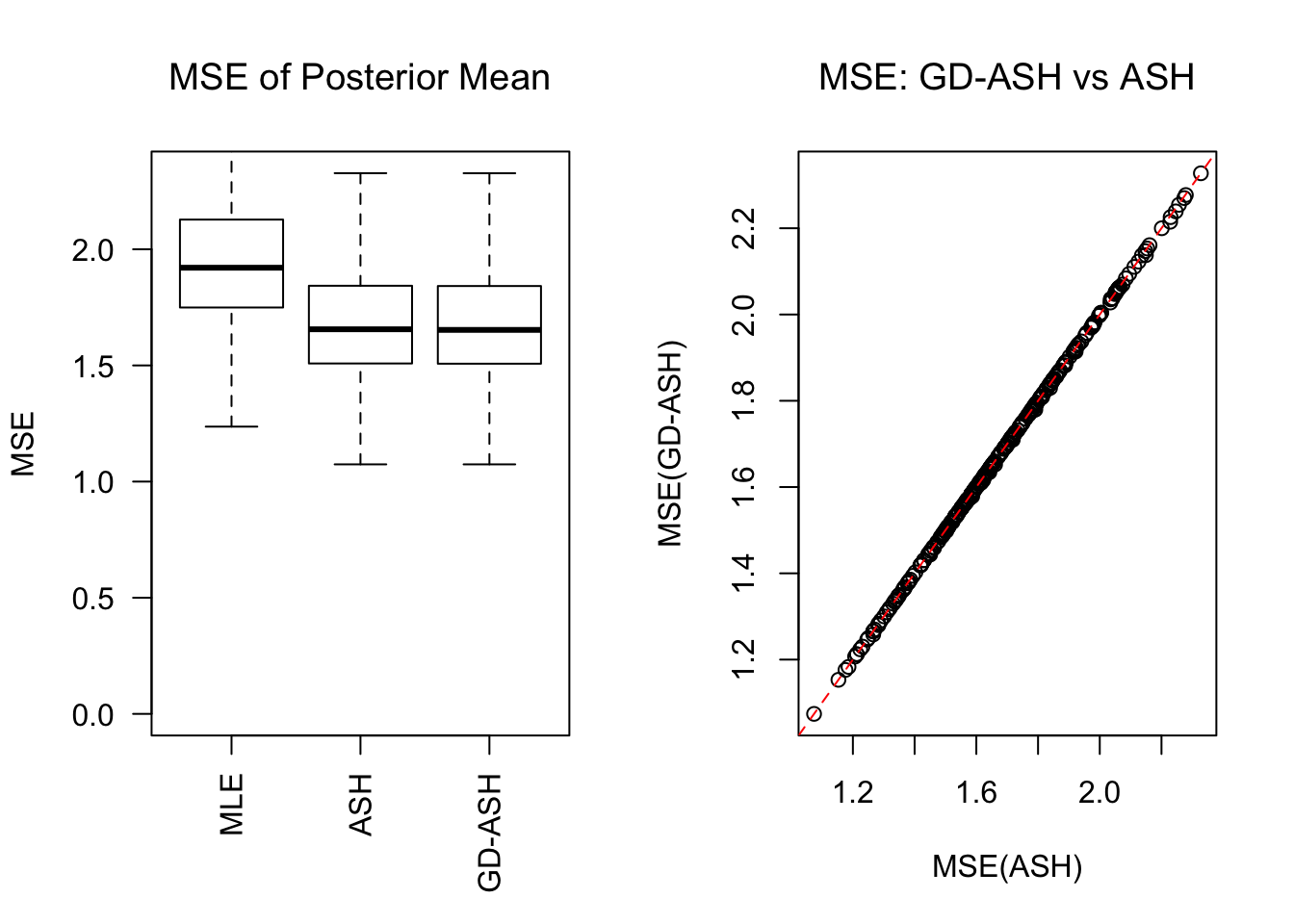
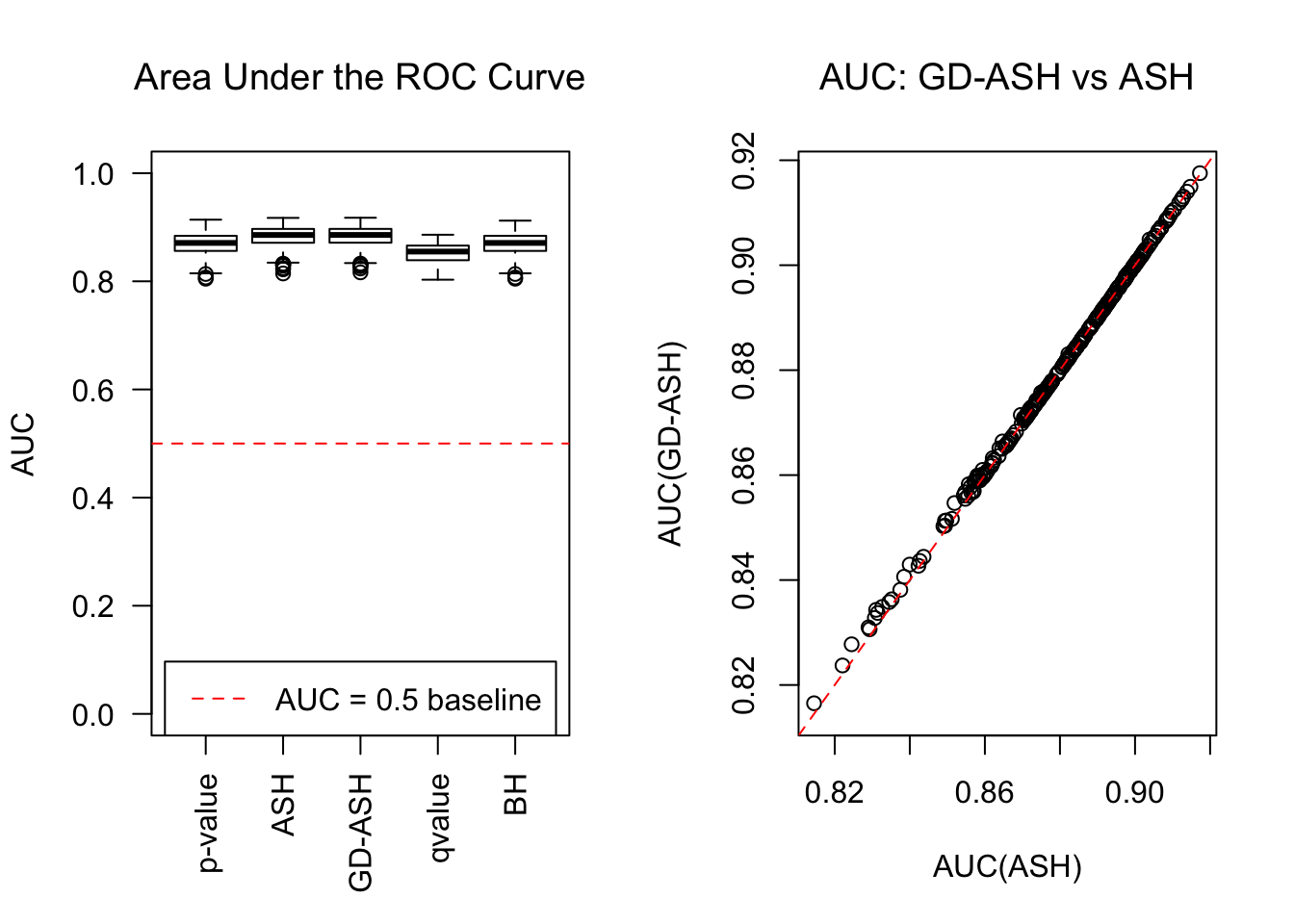
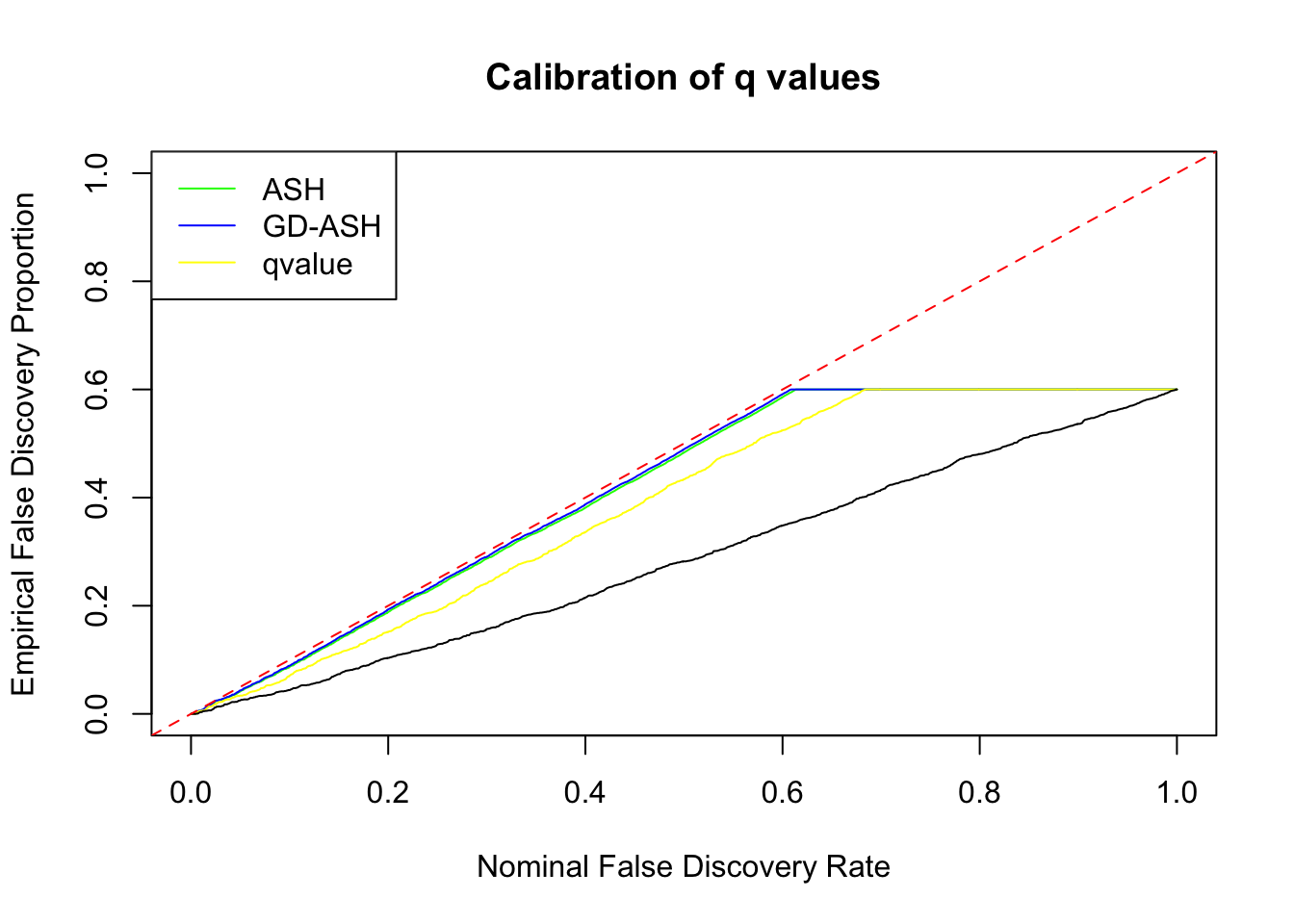
Session information
sessionInfo()R version 3.3.3 (2017-03-06)
Platform: x86_64-apple-darwin13.4.0 (64-bit)
Running under: macOS Sierra 10.12.5
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
loaded via a namespace (and not attached):
[1] backports_1.0.5 magrittr_1.5 rprojroot_1.2 tools_3.3.3
[5] htmltools_0.3.6 yaml_2.1.14 Rcpp_0.12.10 stringi_1.1.2
[9] rmarkdown_1.5 knitr_1.15.1 git2r_0.18.0 stringr_1.2.0
[13] digest_0.6.12 evaluate_0.10 This R Markdown site was created with workflowr