CASH Simulation, Part 5: pFDP Variability
Lei Sun
2017-11-15
Warning in as.POSIXlt.POSIXct(Sys.time()): unknown timezone 'zone/tz/2017c.
1.0/zoneinfo/America/Chicago'Last updated: 2017-11-22
Code version: beffb01
Introduction
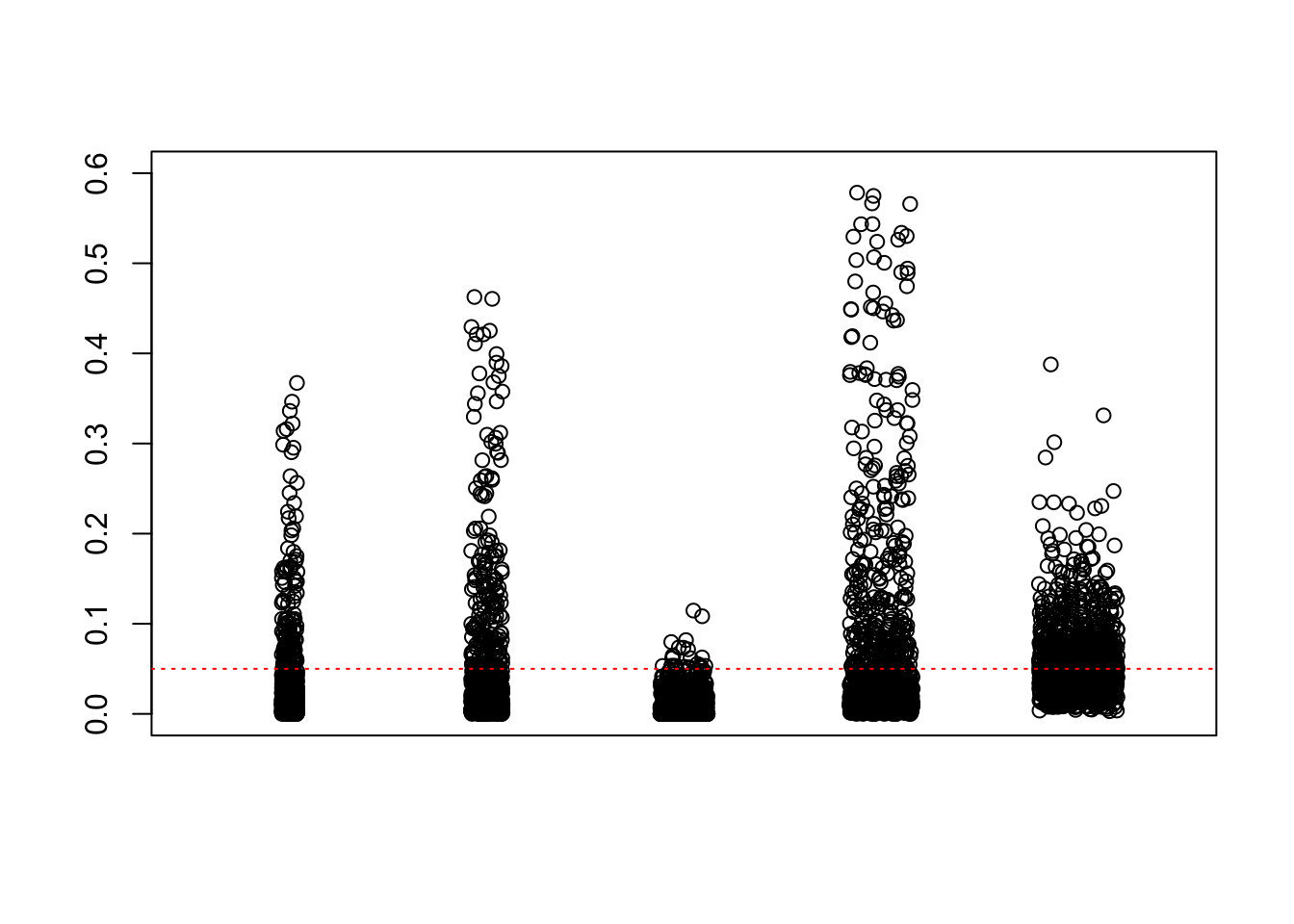
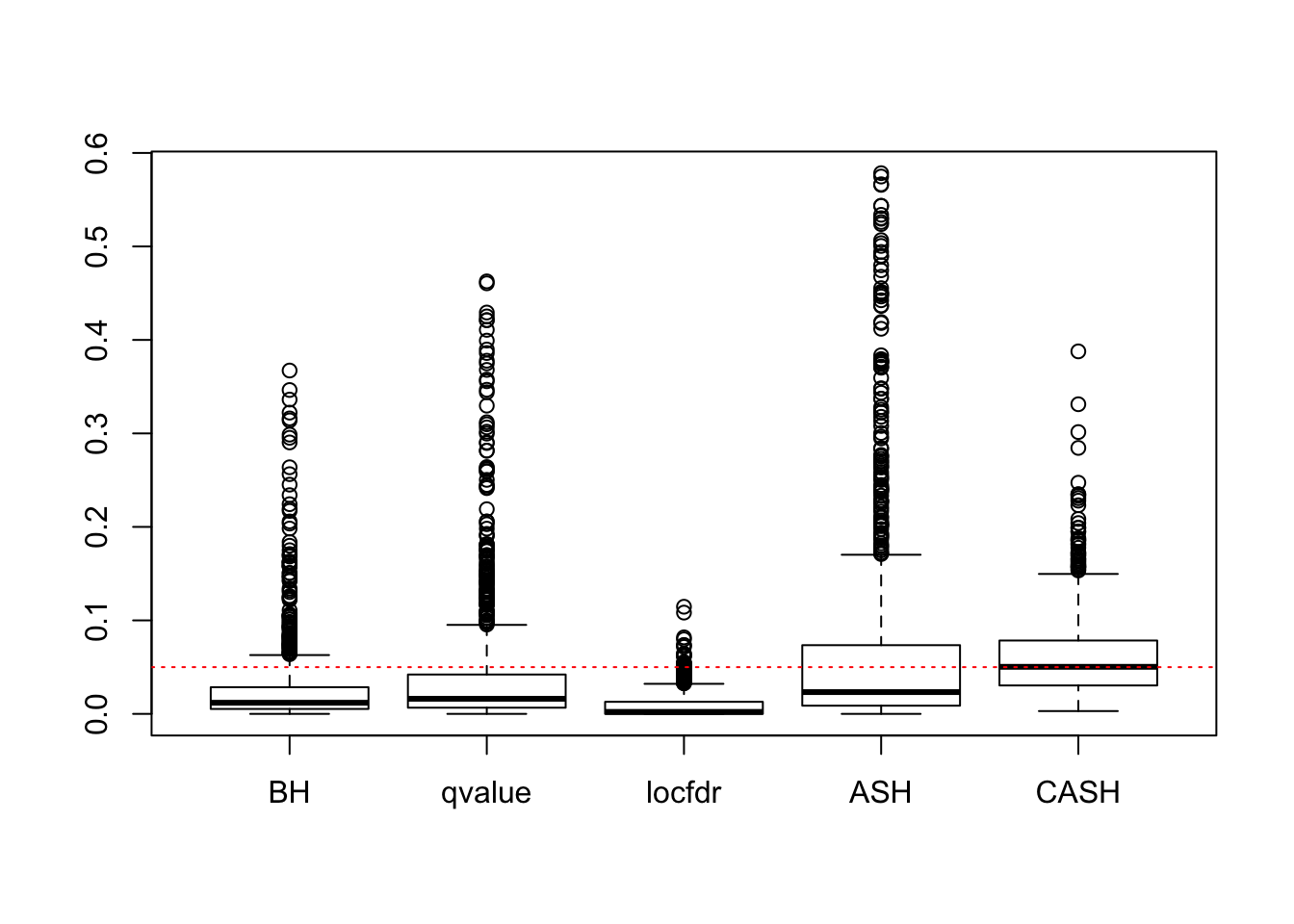
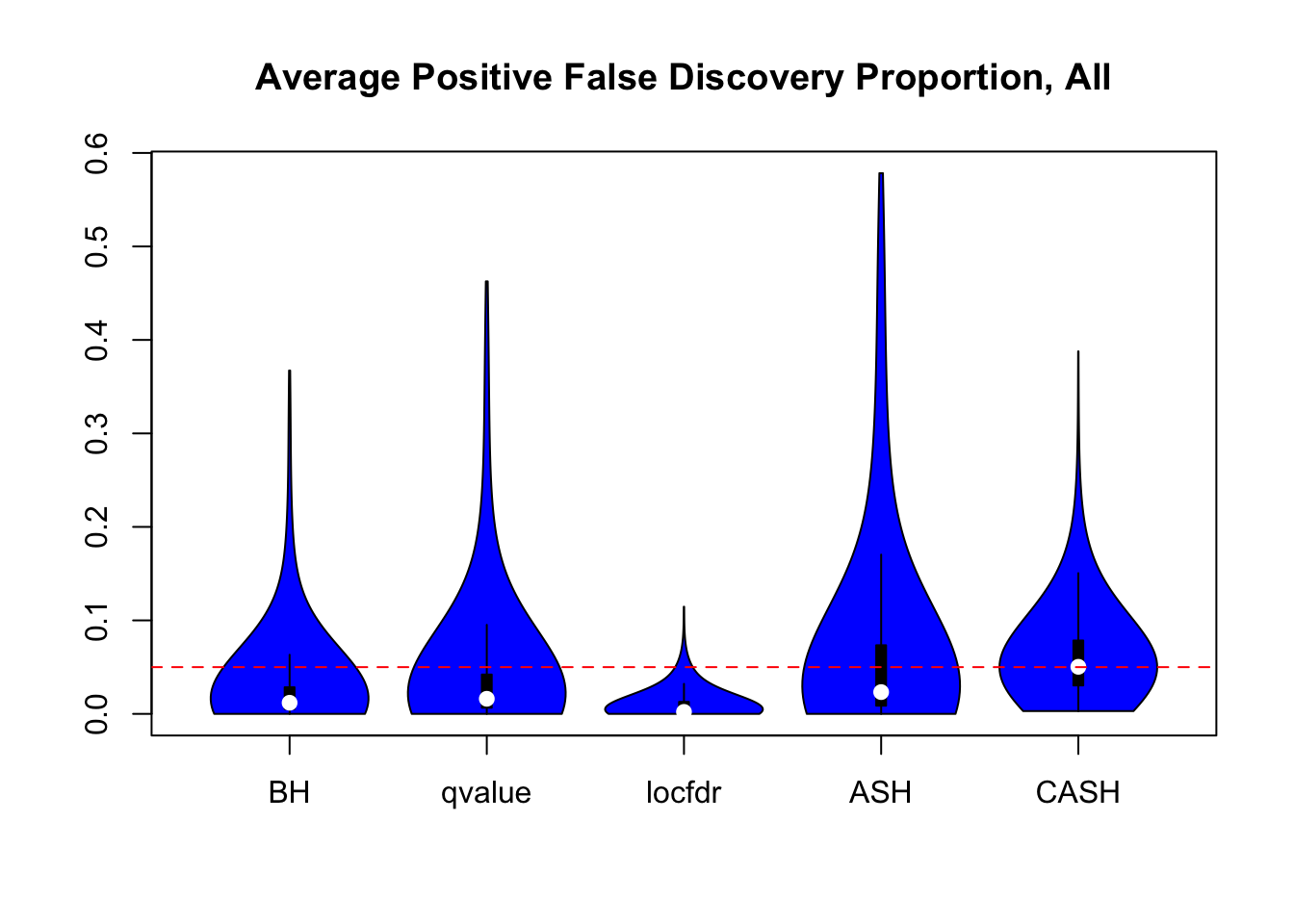
We are using the data obtained from the large-scale simulation with real data to show the variability of the positive false discovery proportions obtained by different methods at nominal \(q\) value $ = 0.05$. The true effect distribution is \(0.6\delta_0 + 0.3N(0, 1) + 0.1N(0, 2^2)\).
z.mat <- readRDS("../output/z_null_liver_777.rds")
se.mat <- readRDS("../output/sebetahat_null_liver_777.rds")
beta.list <- readRDS("../output/beta.list.rds")
qvalue.list <- readRDS("../output/qvalue.list.rds")
qvalue.BH <- qvalue.list$qvalue.BH
qvalue.qvalue <- qvalue.list$qvalue.qvalue
qvalue.locfdr <- qvalue.list$qvalue.locfdr
qvalue.ash <- qvalue.list$qvalue.ash
qvalue.gdash <- qvalue.list$qvalue.gdash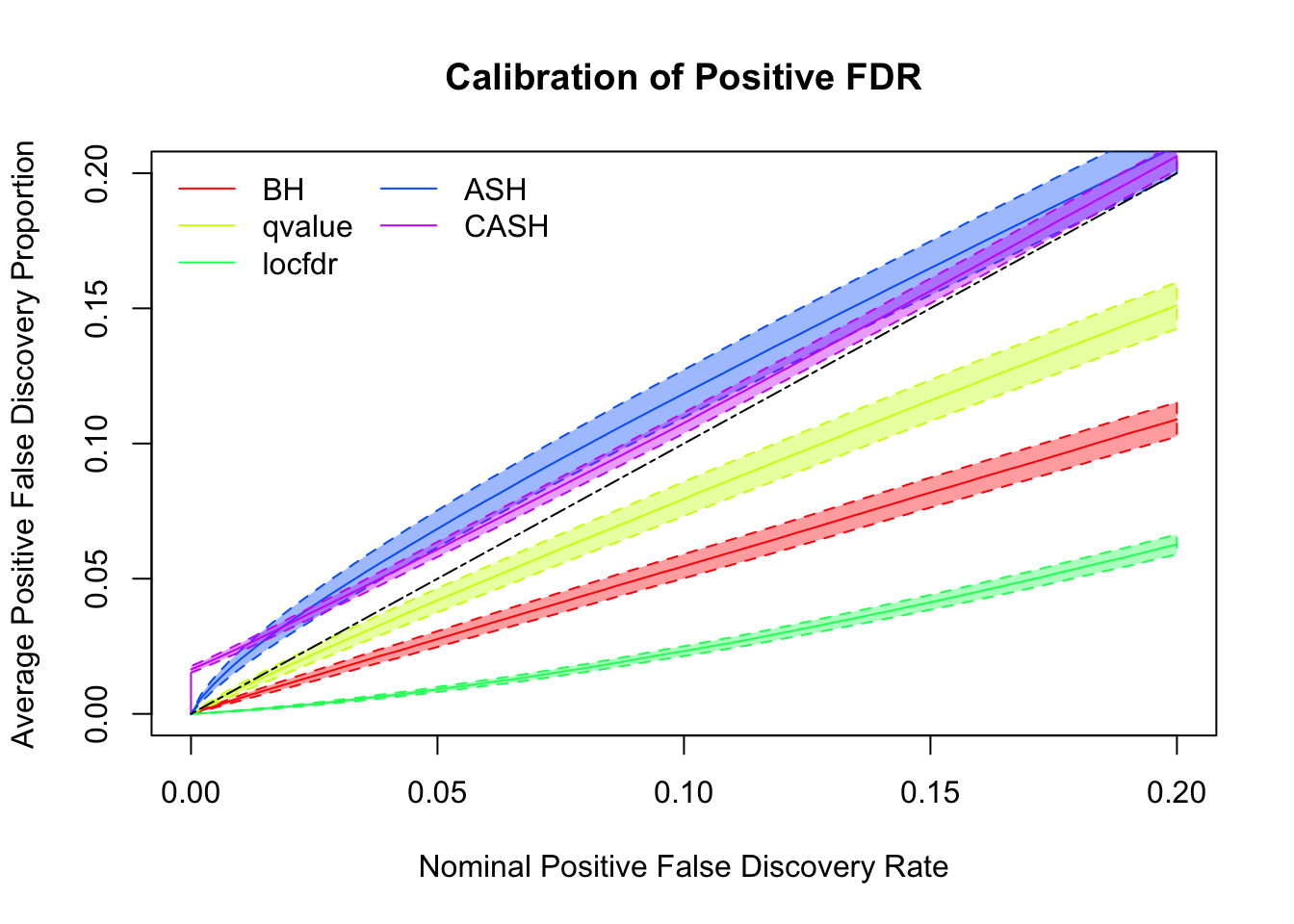
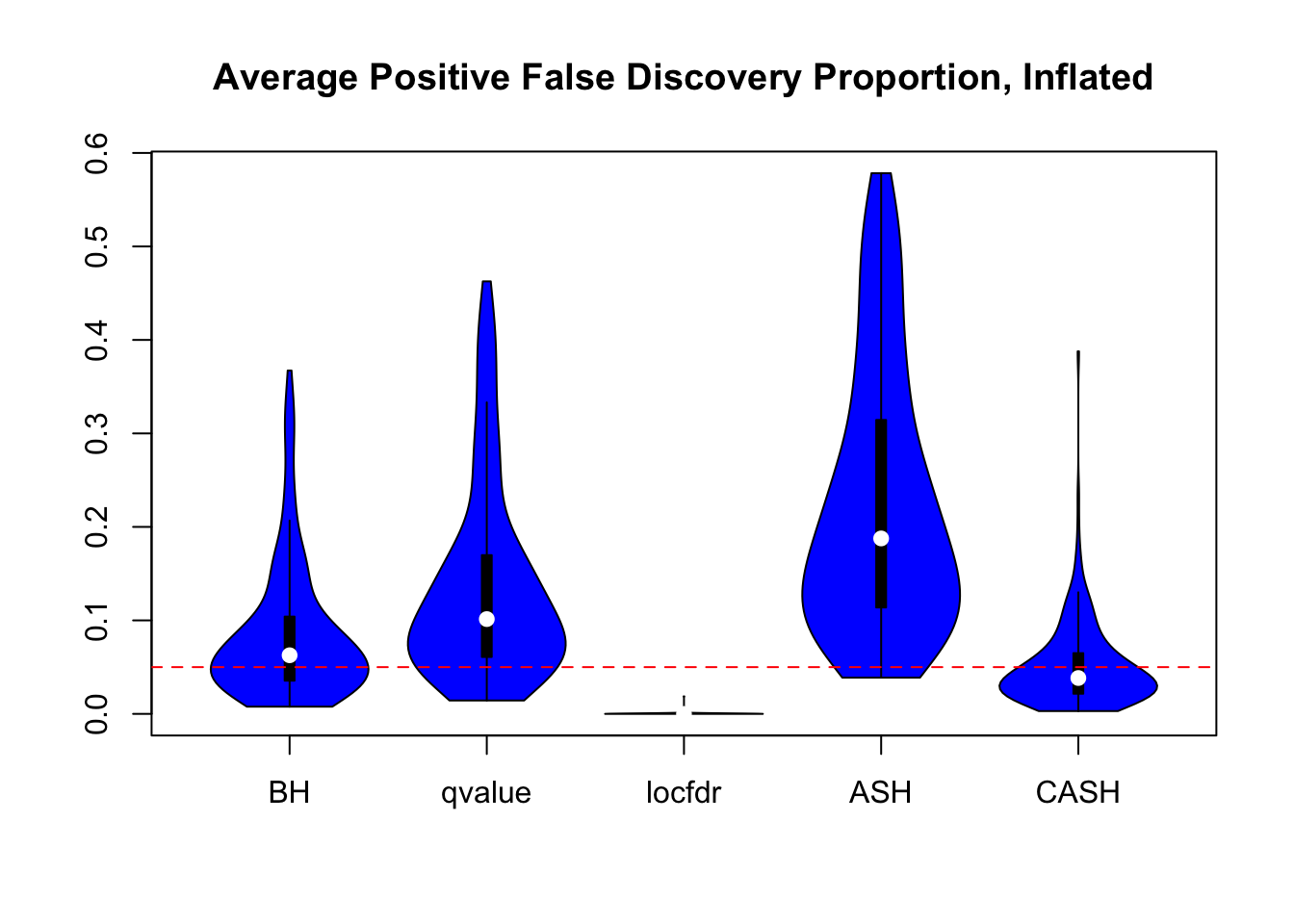
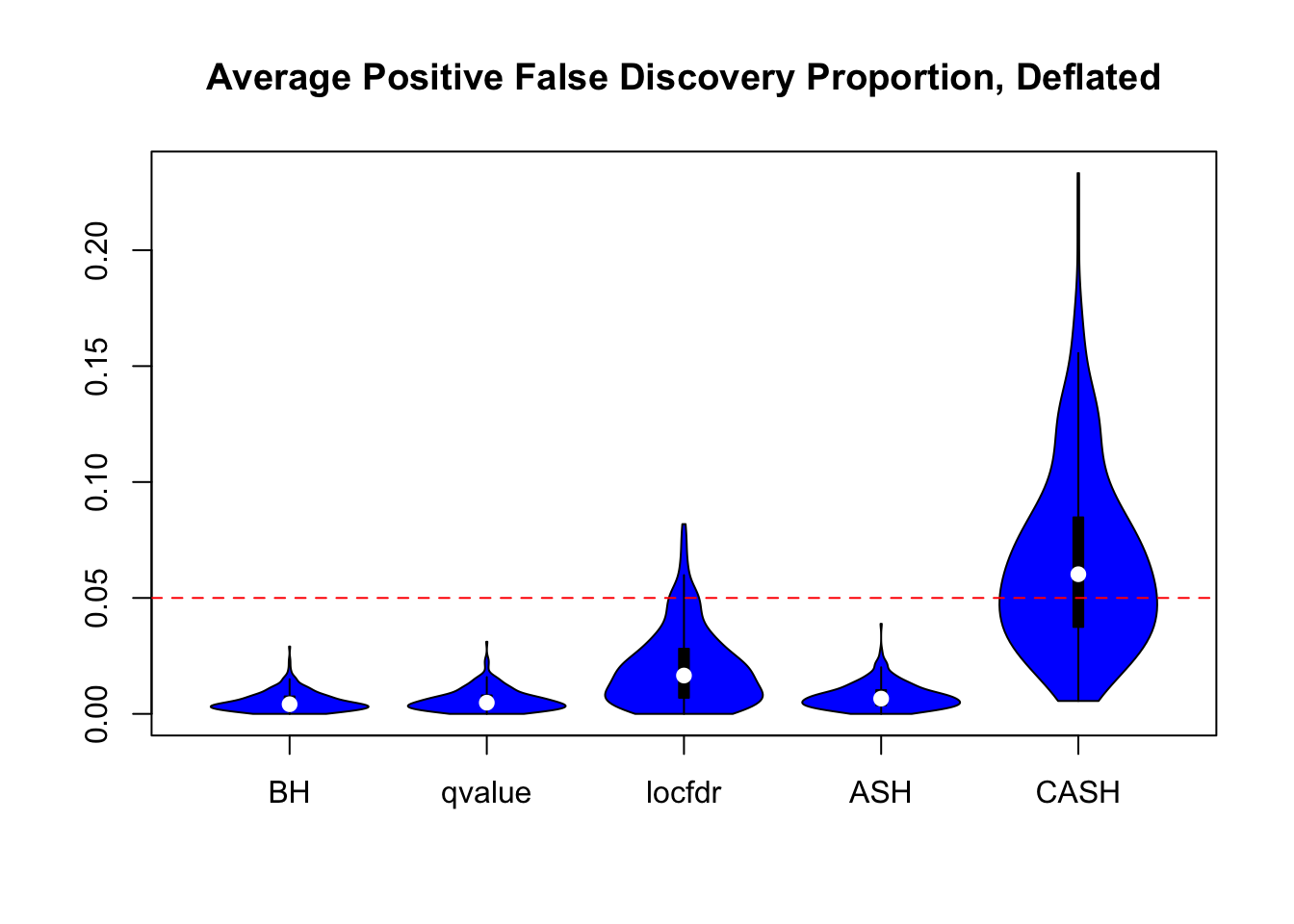
Session information
sessionInfo()R version 3.4.2 (2017-09-28)
Platform: x86_64-apple-darwin15.6.0 (64-bit)
Running under: macOS High Sierra 10.13.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/3.4/Resources/lib/libRlapack.dylib
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] vioplot_0.2 sm_2.2-5.4
loaded via a namespace (and not attached):
[1] compiler_3.4.2 backports_1.1.1 magrittr_1.5 rprojroot_1.2
[5] tools_3.4.2 htmltools_0.3.6 yaml_2.1.14 Rcpp_0.12.13
[9] stringi_1.1.5 rmarkdown_1.6 knitr_1.17 git2r_0.19.0
[13] stringr_1.2.0 digest_0.6.12 evaluate_0.10.1This R Markdown site was created with workflowr